Use of Buckwheat Seed Protease Inhibitor Gene for Improvement of Tobacco and Potato Plant Resistance to Biotic Stress
N. V. Khadeeva1, E. Z. Kochieva2,3, M. Yu. Tcherednitchenko3, E. Yu. Yakovleva1, K. V. Sydoruk4, V. G. Bogush4, Y. E. Dunaevsky5, and M. A. Belozersky5*
1Vavilov Institute of General Genetics, Russian Academy of Sciences, ul. Gubkina 3, 119991 Moscow, Russia2Bioengineering Center, Russian Academy of Sciences, pr. 60-letiya Oktyabrya 7/1, 117312 Moscow, Russia
3Russian State Agricultural University, Timiryazev Moscow Agricultural Academy, ul. Timiryazevskaya 49, 127550 Moscow, Russia
4FSUE State Scientific Center of Russian Federation “State Scientific Research Institute of Genetics”, 1-yi Dorozhnyi Proezd 1, 117545 Moscow, Russia
5Belozersky Institute of Physico-Chemical Biology, Lomonosov Moscow State University, 119992 Moscow, Russia; fax: (495) 939-3181; E-mail: mbeloz@belozersky.msu.ru
* To whom correspondence should be addressed.
Received July 4, 2008; Revision received September 29, 2008
The possibility to use agrobacterial transformation of leaf discs to produce resistance to bacterial infections in tobacco and potato plants by introduction of a single gene encoding the serine proteinase inhibitor BWI-1a (ISP) from buckwheat seeds is shown. All studied PCR-positive transgenic plants exhibited antibacterial activity in biotests. It was shown that the presence of just a single gene of serine proteinase inhibitor provides sufficient protection at least against two bacterial phytopathogens, Pseudomonas syringae pv. tomato and Clavibacter michiganensis sbsp. michiganensis. The biotest including tobacco plant infection by the white wings butterfly in the green house has also demonstrated the existence of protective effect in transgenic tobacco plants. Significant genotypic variations in the protection efficiency were found between members of different genera of the same family (potato and tobacco) as well as between different lines of the same species. Northern blot analysis of four transgenic potato lines and three tobacco lines transformed by a vector plasmid containing the ISP gene of serine proteinases BWI-1a from buckwheat seeds has shown the presence of the expected size mRNA transcript.
KEY WORDS: agrobacterial transformation, serine proteinase inhibitor gene, tobacco, potato, protection against phytopathogensDOI: 10.1134/S0006297909030031
Abbreviations: 6BAP, 6-benzylaminopurine; ISP, gene encoding the serine proteinase inhibitor BWI-1a from buckwheat seeds.
During evolution, plants elaborated a number of efficient mechanisms
providing for their protection against phytopathogens and insect pests.
Among these mechanisms the use of different plant proteins exhibiting
toxic or antimetabolic activities towards pathogens are of great
interest. These proteins include hydrolase inhibitors, lectins,
ribosome-inactivating proteins, enzymes, etc. All of these can be to
some extent involved in the creation of a protective barrier at early
stages of infection. Proteinase inhibitors are also among protective
proteins because both insect pests and pathogenic microorganisms use
proteolytic enzymes for penetration into the host plant tissues. Plants
counteract this by synthesis of constitutive and induced proteinase
inhibitors. Most of studied inhibitors are inactive towards their own
proteinases of seeds, but rather efficiently inhibit proteolytic
enzymes of animal and bacterial origin as well as proteinases of the
alimentary canal of insect pests [1].
Independent researchers repeatedly showed the possibility to use genes of these inhibitors in gene-engineering works for obtaining plants resistant to insect pests. It was shown that introduction into tobacco seeds of trypsin inhibitor from Vigna (cowpea) significantly increased plant resistance to tortrix moth [2]. A number of the pathogen resistance genes isolated from different plants exhibited protective effect in transgenic tobacco plants [3, 4]. Potato and soybean proteinase inhibitors introduced into tobacco and potato plants increased resistance to nematodes [5-7]. It is supposed that besides inhibition of the alimentary canal enzymes, proteinase inhibitors may disturb in insects metabolic processes activated by proteolytic enzymes. There are data allowing one to judge about the ability of some proteinase inhibitors to provide for plant protection not only against pests, but against pathogenic microflora as well [8].
In some recent works, two or three genes whose products have different mechanisms of action have been used simultaneously to enhance resistance to pathogens and pests [8-11]. Thus, expression of two different protective genes, tobacco β-1,3-glucanase (GLU) and lucerne defensin (alfAFP), in transgenic tomato plants and their seed progeny T1 enhanced resistance to the bacterium Ralstonia solanacearum. In this case, from one to three gene copies were inserted into the plant genome, and the resistance level approximately corresponded to their expression level with observance of the gene dose [12]. Besides, synergism of protective gene expression and the promising character of such approach to obtaining phytogen-resistant plant cultures have been shown in a number of works [13]. The possibility of obtaining tobacco plants resistant to microbial and fungal pathogens was shown [10] by introduction of genes of the precursor of the Nicotiana alata and barley β-chordothionine proteinase inhibitor, and it was also shown that both proteins are synthesized in transgenic plants and have a protective function, and this feature is inherited for at least two seed generations. It was shown in numerous works that various plant inhibitors of proteolytic enzymes, including serine protease inhibitors from barley and maize, inhibit growth of many phytopathogenic fungi and, in addition, there are data showing the use of these inhibitors for protection against some bacteria [14].
However, it should be taken into account that in the case of combined coexistence, pathogenic microorganisms and plant pests become adapted to life on the hosts containing proteinase inhibitors. In this case they are able either to synthesize new proteinases, insensitive to the exogenous inhibitor, which even sometimes belong to a different class or family of proteolytic enzymes (for example, to cysteine instead of serine or chymotrypsin and elastase instead of trypsin) [15-18] or to cleave inhibitors by other proteinases insensitive to the exogenous inhibitor [19-21]. Because of this, a continuous search for new inhibitors from unrelated plants, capable of opposing such pathogen adaptation, is under way. In this respect, the proteinase inhibitor BWI-1a from buckwheat seeds may be rather promising, because on one side it quite efficiently inhibits activity of serine proteases that are the main proteolytic enzymes in many microorganisms and insect pests, and on the other side it exhibits biological activity inhibiting growth and germination of pathogenic microflora [22].
The BWI-1a inhibitor was isolated earlier from seeds of buckwheat Fagopyrum esculentum. It is a component of the group of trypsin anionic inhibitors from these seeds, which inhibit growth of Alternaria alternata and Fusarium oxysporum mycelia and proteinases of animal and bacterial origin [23]. Although the role of these proteinase inhibitors in the host plant protective reactions is still not clear, data on inhibition of bacterial enzymes by these proteins point to their possible participation in plant protection against bacterial infection. Chemical synthesis of the gene encoding the proteinase anionic inhibitor BWI-1a of buckwheat was carried out, the structure of which was designed on the basis of amino acid sequence of the protein. The resulting gene was cloned and translated in a cell-free system of rabbit reticulocytes [24]. Translation resulted in a recombinant product corresponding by molecular mass to the proteinase inhibitor BWI-1a.
At the same time, the mechanisms of plant interactions with phytopathogenic microorganisms, the dependence of the extent of the protective effect on the plant species and the introduced gene dose and origin, the extent of expression and full value level of the target protein-inhibitor, as well as the possibilities of preservation of the acquired resistance in the progeny still remain poorly studied. These problems become especially relevant in connection with recent achievements in biotechnology and appearance of numerous genetically modified plants. The integration of such plants into agricultural practice causes pronounced anxiety in the community due to complete ignorance concerning their effects on the environment and health of people.
The aim of this work was to study and characterize the tobacco and potato transgenic plants expressing the gene of the buckwheat serine proteinase anionic inhibitor BW1-1a (ISP) and to study the role of this class of inhibitors in providing for plant protection against pathogens.
MATERIALS AND METHODS
Plant material. Aseptic tobacco plants (Nicotiana tabacum L.) of Samsun variety and potato (Solanum tuberosum L.) of Reserve variety were used in this work. The aseptic tobacco plants and regenerants were maintained on phytohormone-free Gamborg’s B5 medium [25], aseptic potato plants were kept on MS (Murashige–Skoog) medium [26] with addition of 0.5 µg/ml 6-benzylaminopurine (6BAP). Cultures were grown under 16 h photoperiod at 23-26°C.
Transforming vector. Transformation was achieved using Agrobacterium tumefaciens A281 strain carrying a vector construct on the basis of plasmid pBI 101 with inserted 213 bp ISP gene under a strong constitutive 35S promoter of the cauliflower mosaic virus and the neomycin phosphotransferase gene nptII as a selective marker. The target gene structure was designed on the basis of amino acid sequence of the proteinase anionic inhibitor BWI-1a isolated from buckwheat seeds. The ISP gene was obtained by assembly of a 213-member DNA duplex using PCR by cascade joining the extended polynucleotides obtained by direct chemical synthesis [24, 27]. Bacterial culture for transformation was grown overnight in liquid LB medium with 100 µg/ml kanamycin, then plant explants were diluted 4-5 times for incubation.
Plant transformation. Plants were transformed according to Horsch et al. [28] with addition of 400 µg/ml cysteine to media for co-cultivation and regeneration [29]. Primary regenerants were obtained from transformed leaf segments on the following regeneration media: for tobacco 2 µg/ml 6BAP and 0.2 µg/ml naphthylacetate; for potato 3 µg/ml 6BAP was added to the medium. For transformant selection, kanamycin sulfate (100-150 µg/ml) was added, and 500-250 µg cefotaxim was added to remove agrobacteria. In addition, the repeated transformation of leaf discs of the tobacco transgenic line C18 was carried out and a single secondary transformant (IItr) was selected.
Analysis of transformed plants. Total plant DNA was isolated as described in [30]. PCR analysis was carried out according to standard techniques using primers complementary to the gene nptII sequences. The PCR mixture containing DNA vector (~20 pg) was used as positive control and the PCR mixture without DNA served as negative control. Southern blotting was done according to the standard technique [31]. A DNA fragment complementary to the ISP gene was used as a probe. For Northern blotting, poly(A)+-mRNA was isolated using a Quickprep micro mRNA purification kit (USA). The 1.2 kb fragment of vector construct containing the ISP gene sequence was used as a probe, and the 1.8 kb long DNA probe containing the Arabidopsis α-tubulin gene TUA3 was used as control in Northern blotting. To visualize the Southern and Northern blottings, the Phosphorimager system (Kodak, USA) was used.
Biotests. For determination of antibacterial activity of control and transgenic plants, pulverized sprout (stems and leaves) fragments were placed in wells on dishes with agarized LB medium inoculated in advance with bacterial cultures grown for 16-18 h in a liquid LB medium. A model culture (E. coli strain XL1 blue), a strain of the tomato bacterial mottling pathogen Pseudomonas syringae pv. tomato, and a strain of the tomato bacterial cancer pathogen Clavibacter michiganensis subsp. michiganensis were obtained through the courtesy of the Department of Plant Protection of the Russian State Agricultural University, Timiryazev Moscow Agricultural Academy, Russian Academy of Agricultural Sciences. Cultures were incubated at 23-25°C and the extent of antibacterial effect was estimated after 24 h incubation. Homogenates of non-transformed tissues and of those transformed by the ISP gene-free vector were used as controls.
Fungicidal activity was analyzed on Petri dishes with hormone-free MS medium [26] inoculated with spores of the fungus Fusarium sp. Two days after fungus accumulation, the ground sprout (stems and leaves) fragments of control and transgenic plants were applied onto a dish and incubated at room temperature (22-25°C). Results of the test were estimated 48-72 h later. Tissues of non-transformed tobacco and potato lines were used as control. The number of biological repetitions varied from 5 to 16. Standard statistical data processing determined the mean standard arithmetic deviation.
Determination of inhibitor activity. Tobacco roots and seeds were ground in liquid nitrogen, and the inhibitor was extracted with 0.1 M phosphate buffer, pH 6.8, for 2 h at 4˚C. Trypsin solution (5 µl, 1 mg/ml) was added to 1-10 µl extracted inhibitor, and the mixture was incubated at room temperature for 10 min. Residual trypsin activity was determined according to Erlanger et al. [32] using N-benzoyl-DL-arginine p-nitroanilide (0.2 mM) in 0.1 M phosphate buffer, pH 7.0, as substrate. The inhibitor amount lowering the solution optical density by 0.01 under conditions of trypsin activity determination was taken as the activity unit.
RESULTS AND DISCUSSION
Obtaining and analysis of transformed plants. The use for plant transformation of the technique of Horsch et al. [28] modified by Olhoft and Somers [29] (using cysteine) made it possible to increase the transformation frequency. The variant in which both co-cultivation and regeneration were carried out in the presence of cysteine appeared to be the best. It should be noted that cysteine in the used concentration (400 µg/ml) inhibited agrobacterial growth, which increased the passage duration and lowered the number of passages in the presence of antibiotics. It is also possible that this contributed to a higher yield of plants-regenerants.
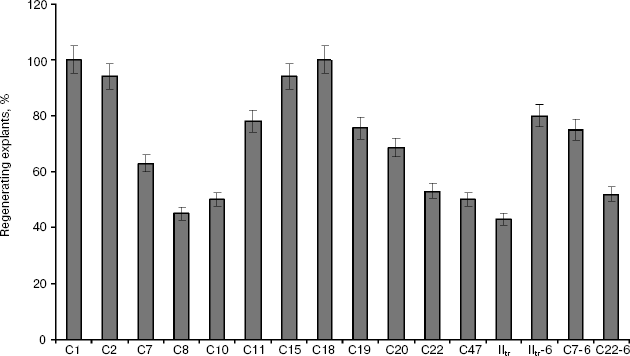
All primary transformants formed roots in the presence of kanamycin, and therefore as an additional checkup the sensitivity to kanamycin of transformant callus- and morphogenesis (i.e. de- and re-differentiation) was studied. The explants of all studied lines formed callus and roots on media for callus genesis and also retained the ability to form adventitious sprouts on regeneration media with kanamycin up to concentration of 400 µg/ml, although different lines somewhat differed from each other by intensiveness. Besides, the lines retained the ability to grow and regenerate in the presence of kanamycin after 1.5-year passage and regeneration on the kanamycin-free medium (Fig. 1), whereas callus- and morphogenesis in control plant was observed only on media free of selective agent.
Individual tobacco and potato lines, transformed by the vector construct carrying the ISP gene, were analyzed by PCR in the presence of cysteine. In the case of tobacco, positive results in PCR analysis were obtained for 90.5% of regenerants and in the case of potato for 60%. When the usual method of Horsch et al. was used, the transformation frequency for the potato leaf segments was 43% and for tobacco about 53%. Thus, in the case of co-cultivation and regeneration in the presence of cysteine transformation frequency and efficiency significantly increased for both plant species. High percentage of transgenic lines obtained upon tobacco transformation can be explained by the chosen scheme of selection in the presence of kanamycin at high concentration (150 µg/ml), which drastically reduced the number of regenerated pseudo-transformants. It appeared that for potato, selection at so high kanamycin concentrations is impossible due to complete inhibition of growth. Thus, the use of this technique made it possible to significantly increase both the yield of regenerants on the whole and the fraction of transformed plants, probably due to lowered necrotization of damaged tissues and increased stable transfer of T-DNA into plant cells.Fig. 1. Regeneration intensiveness in leaf explants of primary transgenic tobacco lines (C1-C47) in the presence of kanamycin and in the same lines after 1.5-year-long reiterated regeneration in non-selective conditions (IItr-6, C7-6, C22-6).
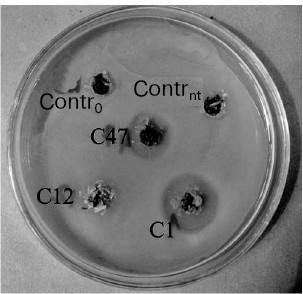
Biotests. Antibacterial activity of transgenic lines was determined using dish tests. Tissues of non-transformed control tobacco plant (Contrnt) and that transformed by the ISP gene-free vector (Contr0) practically did not inhibit growth of any used bacterium, whereas tissues of all analyzed transgenic plants exhibited antibacterial activity (the table). Evidently, transformants synthesized a target functional protein exhibiting inhibitory effect on bacterial growth. However, significant differences were detected in the extent of inhibition of different bacterial species. It should be noted that selection of bacterial strains C. michiganensis and P. syringae is explained by the fact that they secrete serine proteinases during their vital activities [33, 34]. Transgenic clones inhibited growth of P. syringae and to somewhat lower extent growth of C. michiganensis. Besides, secondary growth of separate C. michiganensis colonies was always observed in the inhibition zone. To the least extent transgenic clones inhibited growth of E. coli; in this case even not all transgenic lines exhibited inhibitory effect. It should be noted that tissues of C12 line, that lost their resistance to kanamycin after passage for 2 years, also lost the ability to inhibit bacterial growth (Fig. 2 and the table).
Antibacterial activity of tissues of primary tobacco transformantsFig. 2. Antimicrobial activity of tobacco plant tissue homogenates towards Pseudomonas syringae. Contrnt, non-transformed tobacco plant; Contr0, a plant transformed by a vector free of ISP gene; C1, C12, and C47, transgenic lines (C12 lost kanamycin-resistance and antibacterial activity).
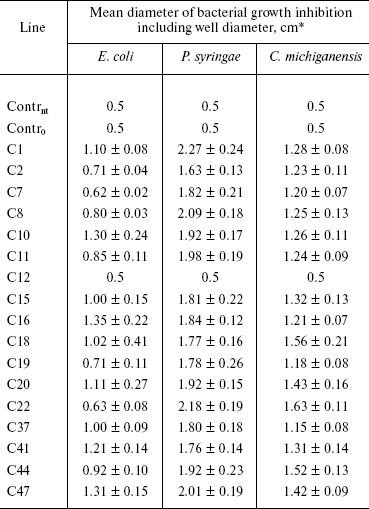
*Well diameter 0.5 cm.
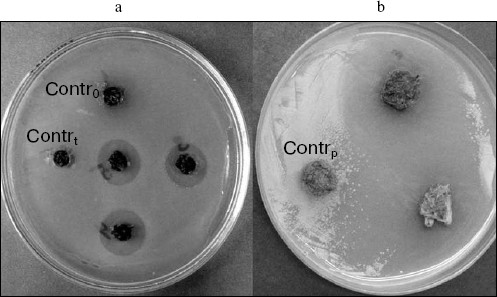
Tissues of transgenic lines of different species highly differed in the extent of antibacterial activity. Thus, wells containing tissue homogenates of transgenic tobacco plants on the lawn of E. coli model culture were surrounded by a halo of growth inhibition of 1.5-1.7 cm in diameter (including the well diameter). In potato, protective effect was much more pronounced in the case of application of the same amount of homogenized tissue, and the halo diameter reached 3-5 cm (Fig. 3). Probably, so pronounced difference in the growth inhibition zone dimensions is caused by individual biological peculiarities of each plant, and possibly by different copy numbers of vector constructs inserted into their genomes.
Analysis of presence and expression of functional ISP gene in cells of potato and tobacco transformed lines. Southern blot analysis was carried out for several transgenic plant lines in which the insert was identified by PCR. The vector construct fragment incorporating the target ISP gene was used as a probe. The common fragment of about 0.6 kb, corresponding to the gene construct fragment, was present in all lines, i.e. insertions in all PCR-positive lines were also identified by Southern blotting. On the whole, single insertions were prevalent [35], which coincides with data from the literature concerning the frequency of gene construct insertions in the case of T-DNA-mediated agrobacterial transformation.Fig. 3. Antibacterial activity of tobacco (a) and potato (b) tissues towards E. coli. Contrt and Contrp, non-transformed tobacco and potato plants, respectively.
Northern blot analysis was carried out for detection of mRNA transcript of the proteinase inhibitor gene and determination of the expression level of this gene in obtained tobacco and potato transgenic lines transformed by the ISP gene-containing construct. Three transgenic lines of potato (Res-24/1, Res-18/12, and Res-17/2) and three tobacco lines (C20, C22, and C12) were selected for analysis of ISP gene expression. According to the above-described biotest, lines Res-24/1, Res-18/12, Res-17/2, C20, and C22 exhibited clear antibacterial activity, whereas line C12 did not reveal such activity (the table) and stopped growing in the presence of kanamycin, which is indicative either of elimination or silencing of this gene.
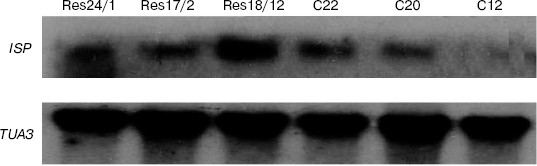
In potato lines Res-24/1, Res-17/2, and Res-18/12 as well as in tobacco lines C22 and C20 the mRNA transcript of expected length of 213 bp was found. As shown in Fig. 4, the ISP gene (of the buckwheat proteinase inhibitor) expression levels differed in different potato and tobacco lines. The highest amount of mRNA was present in the potato line Res-18/12, which correlated with data of biotest. Lower level of ISP gene expression was observed in potato lines Res-17/2, Res-24/1, and in tobacco line C20. However, since all lines tested in this experiment showed antimicrobial activity in biotest, it can be supposed that despite such level of ISP gene expression the formed amount of mRNA was enough to exhibit antibacterial activity of a corresponding protein. Besides, it should be noted that the observed transcription was accompanied by increase in the trypsin-inhibiting activity both in roots (67%) and in seeds (55%) of transformed tobacco lines.
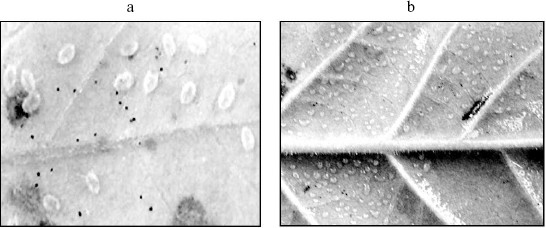
Exposure of tobacco plants by the glasshouse whiteflies (Trialeurodes vaporariorum) observed in the green house has shown that transgenic plants were resistant to the effect of these insects, whereas non-transformed ones were strongly affected by them. Numerous eggs were laid by the whitefly on the control plant leaves and full-value insects emerged from these eggs (Fig. 5a). Only single ovipositions of 1-2 eggs were observed on transgenic plants, and no progeny appeared in this case (Fig. 5b). It should be noted that unlike a number of different insects, serine proteinases are prevalent in the whitefly alimentary canal [36].Fig. 4. Results of Northern blotting of potato (Res-24/1, Res-17/2, and Res-18/12) and tobacco (C20, C22, and C12) transgenic lines. ISP, buckwheat gene of serine proteinase inhibitor; TUA3, Arabidopsis α-tubulin gene used as standard.
Thus, introduction of just a single gene of serine proteinase inhibitor into the plants of heterologous group has shown the possibility of obtaining protective effects against insects and phytopathogenic bacteria. The results indicate that, unlike control plants, those grown by us are able to synthesize certain functional proteins that exhibit protective effect and inhibit development of bacteria. It appeared that the presence of just a single gene of serine proteinase inhibitor provides for sufficient protection against at least two phytopathogenic bacteria. It is quite important to note that to a higher extent transgenic clones inhibited growth of phytopathogenic bacteria P. syringae and C. michiganensis and to a significantly lower extent growth of model culture E. coli, which is probably due to the specificity of proteolytic enzymes of different bacterial species. In addition, significant genotypic distinctions in protection efficiency were observed between members of different genera of the same family (potato and tobacco), which requires further investigation for final clarification of this question. The use of tissue homogenates of the same transgenic tobacco plants for inhibition of growth of phytopathogenic fungi Alternaria alternata and Fusarium culmorum, secreting serine proteinases into the culture medium [37], did not give a significant effect, which might be due to insufficient amount of this gene product for inhibition of these fungal proteinase activities. At the same time, experiments on potato transgenic lines carrying the ISP gene of buckwheat proteinase inhibitor revealed fungicidal effect on Fusarium sp., resulted in cessation of fungus growth at a distance >5 mm from the transgenic line explants, whereas control (non-transformed) variants were completely overgrown [35]. Note that the purified preparation of this protein-inhibitor at concentrations of 0.05-0.5 µg/ml inhibited by 50-100% growth of hyphae of the mycelial fungus A. alternata [38].Fig. 5. Exposure to whiteflies of control tobacco plant (a) and the absence of oviposition on a transgenic plant (b) observed in the greenhouse.
Thus, the possibility of involvement of the recombinant plant inhibitor of proteolytic enzymes in protection of various plants against pathogenic microorganisms and insects is shown in this work. In connection with recent achievements of biotechnology in creation of genetically modified plants characterized by increased resistance to different unfavorable effects, such approach becomes more and more actual, because it not only enables the increase in productivity of cultured plants but contributes to improvement of ecological conditions due to reduced usage of highly toxic protective chemicals.
However, it should be taken into account in the case of creation and investigation of transgenic plants that integration of foreign genes in plants is random; it can proceed in different regions of their genes, both structural and regulatory, and cause alteration of their expression [39, 40]. Results of this work show that in most cases the symptom of kanamycin-resistance is preserved in vegetative and seed progeny of tobacco transgenic plants and during de- and re-differentiation. However, cases of transgene loss or silencing have also been identified and their study will be continued. The analyzed transgenic plants have shown the relationship of the kanamycin-resistance symptom inheritance with the target gene of serine proteinase inhibitor, which confers resistance to pathogens and pests to these plants. Results showing the dependence of the plant protection extent on the genotype, revealed by us upon comparison of transgenic plants of two different genera of the same Solanaceae family, are of special interest. It should also be taken into account that the level of the recombinant protein accumulation in the plant cell depends on many factors, such as the site of inserted gene localization, the used promoter strength, etc. In particular, the use of a strong constitutive promoter, such as the 35S promoter of cauliflower mosaic virus, can result in accumulation of the target product in all plant tissues. The observed differences in the extent of growth inhibition of different bacterial species are also of undoubted interest for further investigation of possible mechanisms of antibacterial activity of transgenic lines expressing this inhibitor.
The authors express their sincere gratitude to students of the Department of Biotechnology (Russian State Agricultural University, Timiryazev Moscow Agricultural Academy) A. Denchik and D. Burdacheva for help in some investigations.
This work was supported in part by the Russian Academy of Sciences Presidium program “Dynamics of Plant, Animal, and Human Gene Pools”, the Russian Foundation for Basic Research (grant No. 09-04-00955), and ISTC (No. 3455).
REFERENCES
1.Dunaevsky, Y. E., Elpidina, E. N., Vinokurov, K.
S., and Belozersky, M. A. (2005) Mol. Biol. (Moscow), 39,
702-708.
2.Hilder, V. A., Gatehouse, A. M. R., Sheerman, S.
E., Barker, R. F., and Boulter, D. (1987) Nature, 330,
160-163.
3.Boulter, D., Edwards, G. A., Gatehouse, A. M. R.,
Gatehouse, J. A., and Hilder, V. A. (1990) Crop Protection,
9, 351-354.
4.Charity, J. A., Bittisnich, D., Anderson, M. A.,
and Higgins, T. J. V. (1999) Mol. Breeding, 5,
357-365.
5.Marchetti, S., Delledonne, M., Fogher, C., Chiaba,
C., Chiesa, F., Savazzini, F., and Giordano, A. (2000) Theor. Appl.
Genet., 101, 519-526.
6.Outchkurov, N. S., de Kogel, J. W., Wiegers, G. V.,
Abramson, M., and Jongsma, M. A. (2004) Plant Biotechnol. J.,
2, 449-458.
7.Vishnudasan, D., Tripati, M. N., Rao, U., and
Khurana, P. (2005) Transgenic Res., 14, 665-675.
8.Datta, K., Baisakh, N., Maung, T., Tu, J., and
Datta, S. K. (2002) Theor. Appl. Genet., 106, 1-8.
9.Abdeen, A., Virgos, A., Olivella, E., Villanueva,
J., Aviles, X., Gabarra, R., and Prat, S. (2005) Plant Mol.
Biol., 57, 189-202.
10.Charity, J. A., Hughes, P., Anderson, M. A.,
Bittisnich, D. J., Whitecross, M., and Higgins, T. J. V. (2005)
Func. Plant Biol., 32, 35-44.
11.Marutasalam, S., Kalpana, K., Kumar, K. K.,
Loganathan, M., Poovannan, K., Raja, J. A., Kokiladevi, E., Samyappan,
R., Sudhakar, D., and Balasubramanian, P. (2007) Plant Cell
Rep., 26, 791-804.
12.Chen, S. Ch., Liu, A. R., and Chzhou, Ts. R.
(2006) Plant Physiol. (Moscow), 53, 756-763.
13.Quyang, B., Li, H.-Y., and Ye, Z.-B. (2003) J.
Plant Physiol. Mol. Biol., 29, 179-184.
14.Valueva, T. A., and Mosolov, V. V. (2002)
Uspekhi Biol. Khim., 42, 193-216.
15.Jongsma, M. A., Bakker, P. L., Peters, J., Bosch,
D., and Stiekema, W. J. (1995) Proc. Natl. Acad. Sci. USA,
92, 8041-8045.
16.Cloutier, C., Jean, C., Fournier, M., Yelle, S.,
and Michaud, D. (2000) Arch. Insect Biochem. Physiol.,
44, 69-81.
17.Wu, Y. R., Liewellyn, D., Mathews, A., and
Dennis, E. S. (1997) Mol. Breeding, 3,
371-380.
18.Bouchard, E., Cloutier, C., and Michaud, D.
(2003) Mol. Ecol., 12, 2439-2446.
19.Girard, C., le Metayer, M., Bonade-Bottino, M.,
Pham-Delegue, M. H., and Jouanin, L. (1998) Insect Biochem.
Mol. Biol., 28, 229-237.
20.Mazumdar-Leighton, S., and Broadway, R. M. (2001)
Insect Biochem. Mol. Biol., 31, 645-657.
21.Zhu-Salzman, K., Koiwa, H., Salzman, R. A.,
Shade, R. E., and Ahn, J.-E. (2003) Insect Mol. Biol.,
12, 135-145.
22.Dunaevsky, Y. E., Gladysheva, I. P., Pavlukova,
E. B., Beliakova, G. A., Gladyshev, D. P., Papisova, A. I., Larionova,
N. I., and Belozersky, M. A. (1997) Physiol. Plant., 101,
483-488.
23.Dunaevsky, Y. E., Pavlukova, E. B., Beliakova, G.
A., and Belozersky, M. A. (1995) Mol. Biol. (Moscow), 28,
1258-1264.
24.Belozersky, M. A., Dunaevsky, Y. E., Drutsa, V.
L., and Dorokhov, Y. L. (1995) Abst. 23rd FEBS Meet., Basel,
Switzerland, p. 210.
25.Gamborg, O. L., Miller, R. A., and Ojima, K.
(1968) Exp. Cell. Res., 50, 150-155.
26.Murashige, T., and Skoog, F. (1962) Physiol.
Plant., 15, 473-477.
27.Belozersky, M. A., Dunaevsky, Y. E., Musolyamov,
A. X., and Egorov, T. A. (1995) FEBS Lett., 371,
264-266.
28.Horsch, R. B., Fry, J. T., Hoffman, N. L.,
Einholtz, D., Rogers, S. G., and Fraley, R. T. (1987) Science,
227, 1229-1230.
29.Olhoft, P. M., and Somers, D. A. (2001) Plant
Cell Rep., 20, 706-711.
30.Edwards, S. K., Johnstone, C., and Thompson, C.
(1991) Nucleic Acids Res., 19, 1349.
31.Maniatis, T., Fritch, E., and Sambrook, J. (1984)
Molecular Cloning [Russian translation], Mir, Moscow.
32.Erlanger, B. F., Kokowsky, N., and Cohen, W.
(1961) Arch. Biochem. Biophys., 95, 271-278.
33.Burger, A., Graffen, I., Engelman, J., Niermann,
E., Pieper, M., Kircher, O., Gertemann, K.-H., and Eichenlaub, R.
(2005) Microbiol. Res., 160, 417-427.
34.Jang, W. H., Kim, E. K., Lee, H. B., Chung, J.
H., and De, O. J. (1996) Biotechnol. Lett., 18,
57-62.
35.Tcherednichenko, M. Yu., Denchik, A. M., and
Kuzmina, N. A. (2004) Abst. IV Conf. of Young Scientists
“Biotechnology in Plant Growing, Animal Husbandry, and
Veterinary”, Moscow, pp. 35-36.
36.Terra, W. R., and Ferreira, C. (1994) Comp.
Biochem. Physiol. B, 109, 1-62.
37.Dunaevsky, Y. E., Matveeva, A. R., Fathkhullina,
G. N., Beliakova, G. A., Kolomiets, T. M., Kovalenko, E. D., and
Belozersky, M. A. (2008) Bioorg. Chem. (Moscow), 34,
317-321.
38.Dunaevsky, Y. E., Pavlukova, E. B., Beliakova, G.
A., Gruban, T. N., Tsybina, T. A., and Belozersky, M. A. (1998) J.
Plant Physiol., 152, 696-702.
39.Ambros, P. F., Matzke, A. J. M., and Matzke, M.
A. (1986) EMBO J., 5, 2073-2077.
40.Feldmann, K. A., Marks, D. A., Christianson, M.
L., and Quatrano, R. S. A. (1989) Science, 243,
1351-1354.