Interaction of Linear Cationic Peptides with Phospholipid Membranes and Polymers of Sialic Acid
A. S. Kuznetsov1,2*, P. V. Dubovskii1, O. V. Vorontsova1, A. V. Feofanov1, and R. G. Efremov1,3
1Shemyakin–Ovchinnikov Institute of Bioorganic Chemistry, Russian Academy of Sciences, ul. Miklukho-Maklaya 16/10, 117997 Moscow, Russia; fax: +7 (495) 336-2000; E-mail: andrej.kuznecov@phystech.edu2Moscow Institute of Physics and Technology, Institutskii Pereulok 9, 141700 Dolgoprudnyi, Moscow Region, Russia
3Higher School of Economics, Myasnitskaya ul. 20, 101000 Moscow, Russia
* To whom correspondence should be addressed.
Received December 27, 2013; Revision received February 11, 2014
Polysialic acid (PSA) is a natural anionic polymer typically occurring on the outer surface of cell membranes. PSA is involved in cell signaling and intermolecular interactions with proteins and peptides. The antimicrobial potential of peptides is usually evaluated in model membranes consisting of lipid bilayers but devoid of either PSA or its analogs. The goal of this work was to investigate the possible effect of PSA on the structure of melittin (Mlt) and latarcins Ltc1K, Ltc2a, and the activity of these peptides with respect to model membranes. These peptides are linear cationic ones derived from the venom of bee (Mlt) and spider (both latarcins). The length of each of the peptides is 26 amino acid residues, and they all have antimicrobial activity. However, they differ with respect to conformational mobility, hydrophobic characteristics, and overall charge. In this work, using circular dichroism spectroscopy, we show that the peptides adopt an α-helical conformation upon interaction with either PSA or phospholipid liposomes formed of either zwitterionic or anionic phospholipids or their mixtures. The extent of helicity depends on the amino acid sequence and properties of the medium. Based on small angle X-ray scattering data and the analysis of the fluorescence spectrum of the Trp residue in Mlt, we conclude that the peptide forms an oligomeric complex consisting of α-helical Mlt and several PSA molecules. Both latarcins, unlike Mlt, the most hydrophobic of the peptides, interact weakly with zwitterionic liposomes. However, they bind anionic liposomes or those composed of anionic/zwitterionic lipid mixtures. Latarcin Ltc1K forms associates on liposomes composed of zwitterionic/anionic lipid mixture. The structure of the peptide associates is either disordered or of β-sheet conformation. In all other cases the studied peptides adopt predominately α-helical conformation. In addition, we demonstrate that PSA inhibits membranolytic activity of Mlt and latarcin Ltc1K. These data suggest that the peptides, due to their high conformational lability, can vary structural and amphiphilic properties in the presence of PSA. As a result, various scenarios of the interaction of the peptides with membranes, whose surface is abundant with anionic polysaccharides, can take place. This can account for difficulties in understanding the structure–functional relationships in interactions of linear cationic peptides with biological membranes.
KEY WORDS: linear cationic peptides, cytolytic peptides, latarcins, melittin, phospholipid liposomes, sialic acid, circular dichroismDOI: 10.1134/S0006297914050101
Abbreviations: CD, circular dichroism; DOPC, dioleoylphosphatidylcholine; DOPE, dioleoylphosphatidylethanolamine; DOPG, dioleoylphosphatidylglycerol; LCP, linear cationic peptide; LPS, lipopolysaccharide; Ltc1K, latarcin 1K from venom of Lachesana tarabaevi spider; Ltc2a, latarcin 2a from venom of Lachesana tarabaevi spider; Mlt, melittin from venom of Apis mellifera honey bee; PSA, polysialic acid; SAXS, small-angle X-ray scattering.
Electrical charge and amphiphilicity are fundamental characteristics of
biomolecules governing their activity in vivo. Amphiphilic
peptides carrying a multitude of electrical charges are physiologically
active due to their ability to interact with receptors and a variety of
biomembrane components [1, 2].
In particular, polycationic peptides are considered as perspective
molecules for antibacterial and anticancer therapy [3, 4]. Such peptides are widely
present in the venoms of insects, arachnids, and snakes [5, 6]. Some of them, e.g.
cardiotoxins, feature compact and highly stable spatial structure due
to the presence of disulfide bonds [7, 8]. In contrast, linear peptides exhibit high
flexibility and are typically able to switch among a number of states
[9, 10]. In this work we
studied amphiphilic peptides with high net positive charge –
linear cationic peptides (LCP).
These include latarcins Ltc1K [11] and Ltc2a [12] isolated from the venom of the Middle Asia Lachesana tarabaevi spider [13, 14] and melittin (Mlt) from the venom of Apis mellifera honey bee [15]. These peptides feature equal length (Table 1), but their net charge varies from +5 (Mlt) to +10 (both latarcins).
Wide-spectrum antibacterial activity and toxicity against cancer cells of different etiology are specific for them [17, 18]. These activities are supposed to arise due to ability of the peptides to form amphiphilic α-helices upon interaction with lipid membranes containing anionic phospholipids [4, 11, 12]. The hydrophobic/hydrophilic properties of such helices are different for various peptides. In some cases this suggests the mechanism of deterioration of the target lipid membrane and suggests an explanation for variation in activities of the peptides toward bacteria of different strains. Alternatively, the effect of LCP on the lipid matrix of the membrane cannot be solely responsible for the whole spectrum of biological activity of the peptides in the course of their interaction with a variety of cells. Indeed, the effect of LCP on Gram-negative microorganisms can be associated with the ability of the peptides to traverse their lipopolysaccharide (LPS) layer. The latter protects the inner plasma membrane of bacteria from outer perturbations [19-21]. For animal cells, polysialic acid (PSA) is one of the polymers that is abundant on the membrane surface [22, 23]. Both LPS and PSA feature periodic distribution of negatively charged moieties. Thus, interaction of LCP with such polymers can modulate the impact on the cell and determine the selectivity of their antibacterial action [24]. Binding of Mlt to heparin, heparan sulfate, and dermatan sulfate, the ubiquitous polyanions of the cell surface, has been characterized through developing a model of their interaction [25]. Such models are lacking for latarcins.
Table 1. Polycationic peptides, used in the
work, and their properties
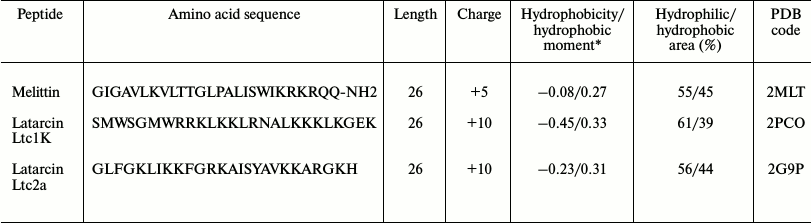
* Value of average hydrophobicity index and hydrophobic moment are given
according to the scale of Eisenberg et al. [16].
The goal of the present work was to study the interaction of LCP (Mlt, Ltc1K, and Ltc2a) with phospholipid membranes in the presence and absence of PSA. The following tasks have been solved. 1) The effect of PSA of different molecular mass on secondary structure of the peptides was evaluated. 2) The possibility of formation of stable associates between these peptides and PSA was investigated. 3) Differences between the three peptides were elucidated with the respect of the influence of PSA on their membranolytic activity. In addition, changes in secondary structure of the peptides in their interaction with liposomes of different lipid composition were investigated. The data cast light on the properties of the peptides determining their interaction with conformationally flexible polyanionic polymers (PSA) and phospholipid membranes.
MATERIALS AND METHODS
Reagents. The molecular mass of the polysialic acid used was either 6.9 or 22.9 kDa. The corresponding substances are abbreviated as PSA-7 and PSA-23, respectively. Both were products of FDS Pharma (Great Britain). Latarcins Ltc1K and Ltc2a were synthesized via the solid-phase technique [13]. The purity of melittin (above 95%) isolated from bee venom was checked via PAGE and gel-permeating chromatography. Poly-L-lysine (15-30 kDa), polyethyleneimine (700 kDa), dioleoylphosphatidylcholine (DOPC), dioleoylphosphatidylethanolamine (DOPE), and dioleoylphosphatidylglycerol (DOPG) were products of Avanti Polar Lipids (USA). Deionized water was obtained with the use of Millipore (France) equipment. Salts were products of Fluka (Switzerland) and fine grade quality of Reakhim (Russia). The following buffers were prepared: PBS (1.7 mM KH2PO4, 5.2 mM Na2HPO4, 150 mM NaCl, pH 7.3), 0.1 M citrate buffer (citric acid-sodium citrate at different ratios, pH 3.2-6.0), and 50 mM HEPES-NaOH, pH 7.0 (Pan Eco, Russia).
Circular dichroism. Circular dichroism (CD) spectra were registered with a J-810 spectropolarimeter (Jasco, Japan) in the 190-250 nm range. The digitization step was 0.2 nm. A cell of 0.1-mm pathlength was used. Aqueous or buffer (PBS, HEPES-NaOH, citrate) solution of the peptides and their mixtures with PSA were used. The concentration of latarcins was 0.3 mM (0.1 mM for Ltc2a in PBS) and of Mlt 0.175 mM. The concentration of PSA-23 was varied from 0.04 to 0.30 mM, and that of PSA-7 from 0.15 to 0.60 mM. For mixtures of peptides with PSA, the spectra of PSA at the appropriate concentration were subtracted. The elements of secondary structure were estimated using the Selcon3, Cdsstr, and Continll programs. The basis for decomposition of the spectra was one with data obtained for as many as 29 globular proteins [26]. The CD spectra of the peptides in the presence of liposomes were acquired at room temperature. The baseline was corrected via subtracting spectra of the liposomes without peptides added. The absorbance value was transformed into ellipticity per amino acid residue using the equation: Θ222 = 100[Θobs]/Nlc, where [Θobs] is ellipticity at 222 nm (mdeg), N is number of residues in the peptide, l is pathlength of the cell (cm) (all spectra of the peptides in the presence of liposomes were obtained with cell of 0.001 cm pathlength), and c is peptide concentration (mM). The concentration of peptides was ~10 µM. The lipid concentration was varied for lipid/peptide ratio (L/P) to be 1 : 60. Buffer of 0.1 M Tris-HCl, pH 7.5, was used. The diameter of liposomes was of 100 nm.
Fluorescence spectra of the tryptophan residue in melittin. The fluorescence spectra were determined in the 290-440 nm range using a Hitachi F4000 fluorimeter (Japan). A 10 × 10-mm cell was used. The slit width for excitation and emission was 5 nm, and the excitation wavelength was 280 nm. Solutions of Mlt in either the presence or absence of PSA were investigated in water, PBS (pH 7.25), 5 mM HEPES-NaOH + 150 mM NaCl (pH 7.0), and 2 M NaCl solution.
Small-angle-X-ray scattering (SAXS). The concentration of aqueous solutions of Mlt was 1.5-4.0 mM and of PSA it was 0.8-8.0 mM. Also, mixtures of peptide–PSA at 1 : 1 and 1 : 2 ratios were used. The SAXS measurements were conducted with the AMUR-K instrument in the A. V. Shubnikov Institute of Crystallography, Russian Academy of Sciences in the laboratory of SAXS. In the spectra obtained a collimation correction was introduced. An order parameter characteristic of the medium heterogeneity was estimated. Distribution of particles over sizes was evaluated for monodisperse systems. The shape of the particles was also determined.
Membranolytic activity of peptides. Liposomes formed of either DOPC or DOPC/DOPG (2 : 1) were used in the experiments. The liposomes were prepared from corresponding stock solutions (10 mM) of the lipid in chloroform–methanol mixture (2 : 1 v/v). An aliquot of the lipid solution was placed in a round-bottom flask and dried for 1 h using a rotary evaporator and then lyophilized for 1 h. The lipid film formed was hydrated with 5 mM HEPES-NaOH buffer in the presence of 100 mM calcein (pH 7.5) for 1 h. Then the film was freeze–thawed five times. Size-calibrated liposomes were obtained using a mini-extruder (Avanti Polar Lipids, USA). The membrane (Whatman, Great Britain) pore size was 200 nm. The liposomes were kept at –4°C. The aliquot of the suspension of the liposomes loaded with calcein was added onto a Sephadex G-50 column (1.1 × 25 cm). The eluent was 5 mM HEPES-NaOH (pH 7.5), and the elution rate was 0.5 ml/min. Fractions of 1 ml were collected. The presence of the liposomes in the fractions was estimated via detection of variation in calcein fluorescence intensity after 10% Triton X-100 solution was added.
The calcein fluorescence was measured with a Hitachi F4000 fluorimeter (Japan). The size of the cell for fluorescence detection was 10 × 10 mm. The width of the slits was of 5 nm, and the excitation wavelength was 495 nm. The fluorescence was detected at 515 nm. The aliquot of the liposome suspension was diluted to 1 ml with 5 mM HEPES-NaOH buffer containing 150 mM NaCl. In the beginning of each experiment, a spontaneous leakage of calcein from liposomes (5-10 min prior to the addition of either peptide or PSA or their mixture) was estimated. The 100%-level of dye escape was obtained after 10 µl of 10% (by weight) Triton X-100 solution was added.
Distribution of hydrophobic–hydrophilic regions on the surface of the peptides. To build maps, the PLATINUM program [27, 28] was used. The conformation of the peptides was assumed to be an ideal α-helix. The models were constructed using the PyMOL program. To calculate values of molecular hydrophobicity potential (MHP) at the surface of α-helices, the set of parameters [29] shifted by +0.03 units of MHP was used. The dependence of MHP upon distance was assumed to be exponential: exp(–r/2). The most hydrophobic regions on the maps were indicated using isolines. The location of Cα-atoms of the amino acid residues was marked.
RESULTS
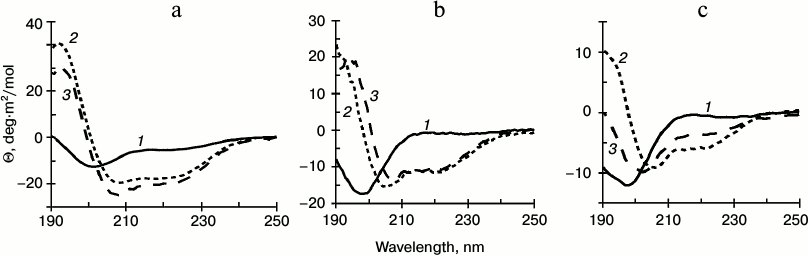
Helix formation in cationic peptides influenced by PSA. Analysis of CD spectra of aqueous solutions of Mlt and latarcins indicated that the peptides are all random-coil in this environment. This agrees with the literature [13, 30]. In all cases, addition of PSA to the peptide solution resulted in an increase of the α-helix contribution in the CD spectrum. In experiments with Mlt, addition of PSA resulted in a substantial (up to 70-80%) increase in α-helix content. This effect did not depend upon PSA concentration in the range of peptide/PSA molar ratios from 8 : 1 to 1 : 1 for PSA-23 and from 2 : 1 to 1 : 2 for PSA-7 (Fig. 1a). Change in pH of the solution from 3 to 7 did not influence the appearance of the spectrum of the Mlt–PSA mixture. Initially, in PBS Mlt displays high α-helix content (about 50%). Addition of PSA-7 at molar ratio 1 : 1 or PSA-23 to molar ratio 2 : 1 resulted in an increase in helix formation, but the effect was weaker compared to the case when the corresponding amounts of PSA were added in water (Fig. 2).
For Ltc1K latarcin, the content of α-helix increased to 60-80% when PSA concentration was increased (Fig. 1b). The maximum effect was observed for Ltc1K/PSA-7 mixture at molar ratio 1 : 1 and for Ltc1K/PSA-23 mixture at ratio 2 : 1. An increase in PSA concentration above these threshold values did not affect the shape of the CD spectrum (data not shown). In PBS (pH 7.3), α-helix formation in Ltc1K in the presence of PSA was found to be substantially lower than in water (Fig. 2).
Fig. 1. CD spectra of Mlt (a) and latarcins Ltc1K (b), Ltc2a (c) in water (1) and in the presence of PSA-7 at ratio 1 : 1 (2) and PSA-23 at ratio 2 : 1 (3). Θ, molar ellipticity.
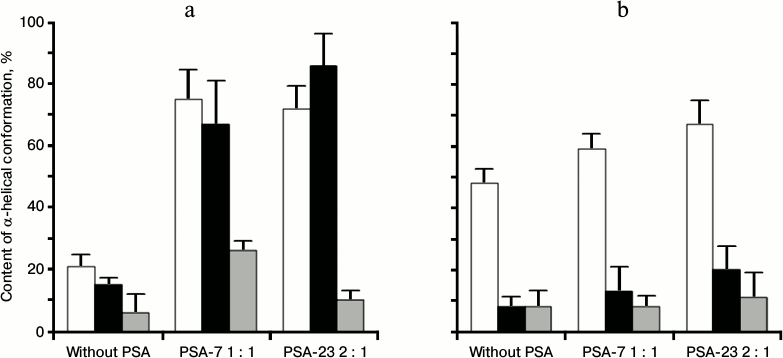
Fig. 2. Helix formation (%, content of α-helical conformation according to CD data) for Mlt (white) and latarcins Ltc1K (black) and Ltc2a (gray) in water (a) and PBS (b) in the presence and absence of PSA.
Addition of PSA-7 to Ltc2a at molar ratio 1 : 1 increased α-helix contribution in the CD spectrum from 7 to 26% (Fig. 2). Addition of PSA-23 at ratio 2 : 1 did not increase α-helix content of this peptide. In all cases Ltc2a showed lower helix content than Ltc1K and Mlt under similar conditions (Fig. 2).
Analysis of the CD spectra of PSA in the pH range 3.3-7.0 at PSA-23 concentration of 0.5-7.2 mg/ml revealed small changes at pH 3.3 only (data not shown). When a distinctly different cationic polymer, polyethyleneimine, was added as a control, no change in the appearance of the CD spectrum of PSA was observed. This was used as a basis for subtraction of the PSA spectra from those of the PSA/peptide mixtures to analyze secondary structure of the peptides.
To evaluate the dependence of the effect of α-helix formation on amino acid sequence of the peptides under the influence of PSA, a number of experiments with another cationic homopolymer, poly-L-lysine (15-30 kDa), were conducted. No changes in the spectra of polypeptides (1.25 mg/ml) after addition of PSA (at ratio poly-L-lysine/PSA 1 : 2 and 1 : 4) were observed (data not shown).
Helix formation in peptides interacting with liposomes. The data on structuring of latarcins in the presence of the liposomes of different composition are given in Table 2. The spectra of all the peptides interacting with zwitterionic liposomes of DOPC (Table 2) were dominated by random conformation. This indicated that only a small portion of the peptide molecules is bound to the liposomes. The ability of the peptides to interact with such liposomes was evidenced by fluorescent dye leakage from these liposomes caused by addition of the peptides [14].
In the case of anionic liposomes of DOPG (Table 2), the α-helix content in Ltc1K is somewhat increased. However, fusion of the liposomes occurs with increase in their size. This was judged by an increase in light scattering, distorting the CD spectra of the peptides.
The most different are the spectra of Ltc1K and Ltc2a in the presence of liposomes of mixed composition, DOPG/DOPE. At varied L/P ratios, Ltc2a bound the liposomes and α-helix formation was detected. Clearly, Ltc1K binds liposomes too, because this peptide interacts with DOPG liposomes readily. However, when DOPE was added, no helix formation occurred in the peptide. Most likely the peptide remains at the surface of the liposomes in a non-helical conformation. It cannot be excluded that the peptide specifically interacts with DOPE. Indeed, if the peptide interacted with DOPG, it became helical. It is important to note that Ltc1K did not cause fluorescent dye leakage from liposomes of this composition [11].
Table 2. Extent of helix formation in
latarcins interacting with liposomes of different composition
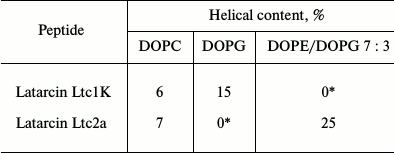
* The peptide precipitates.
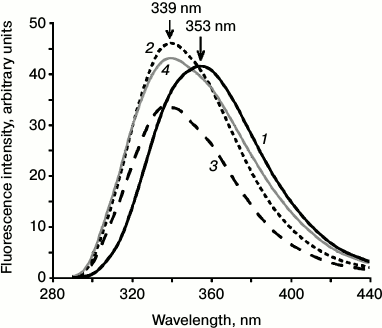
Microenvironment of the Trp residue in Mlt. The fluorescence spectrum of the Trp19 residue in Mlt in water and 5 mM HEPES-NaOH buffer containing 150 mM NaCl exhibited emission maximum at 353 nm. In the presence of 2 M NaCl (conditions of tetramerization of Mlt [15, 31]) in mixtures of Mlt and PSA-7 or PSA-23, a shift of the maximum to shorter wavelength was observed (Fig. 3). This indicates an increase in hydrophobicity of the environment of this residue.
Fig. 3. Emission spectra of fluorescence of the Trp residue of Mlt in water (1) in the presence of PSA of 7 kDa at 1 : 1 (2) and 23 kDa at 1 : 1 (3) and in 2 M NaCl solution (4). The positions of the maxima of fluorescence are indicated with arrows.
Size and shape of complexes of peptides with PSA according to SAXS data. Aqueous solutions of PSA-7 and PSA-23 looked like polydisperse media with an order parameter (a characteristic distance between inhomogeneities of the medium) of ~80 Å. A Mlt/PSA-23 mixture was also polydisperse, with order parameter being ~100 Å. Mlt and PSA-7 mixtures at ratio 1 : 1 and 1 : 2 exhibited a different pattern, corresponding to monodisperse systems. The size of the scattering particles under these conditions was estimated. This system was described most adequately with a model of scattering by spherical particles. Calculation of the distribution of scattering particles over sizes predicted two possible radii for such particles – 3 and 29 Å. The latter was considered as PSA–Mlt complexes, because the size of the complex cannot be less than the size of the peptide itself (see Table 3).
Table 3. Characteristic sizes of melittin
and its oligomeric complexes

* Diameter (D) is given for spherical particles, for
cylindrical-diameter (D) and height (L) of the cylinder.
** Structure from Protein Data Bank (PDB code 2MLT).
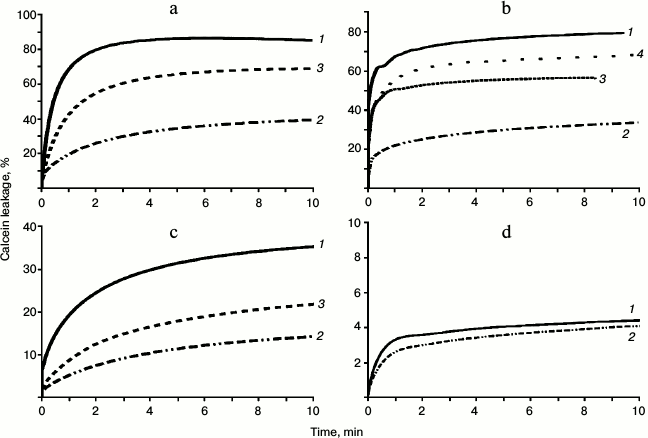
Membranolytic activity of melittin. Melittin showed high membranolytic activity toward liposomes of different composition. For zwitterionic liposomes of DOPC ~70-90% of calcein leakage was observed at the ratio of lipid/Mlt 100 : 1. For the mixed liposomes of DOPC/DOPG (2 : 1) a similar leakage level was observed at lipid/peptide ratio 40 : 1. Activity of Mlt toward DOPC liposomes was found to be ~3 times higher than to the mixed DOPC + DOPG liposomes. A rapid deterioration of the liposomes by Mlt was observed. The rising curves of the fluorescence intensity of calcein reached a plateau at 5-10 min (Fig. 4a).
If the Mlt/PSA complex was formed prior to the addition of Mlt to the liposomes (10-15 min), a substantial decrease in membranolytic activity was observed. PSA-7 induced the strongest inhibiting effect. A decrease in Mlt activity by a factor of 1.5-2.0 was observed even at peptide/PSA ratio 1 : 1. Addition of excess PSA resulted in a stronger decrease in membranolytic activity of Mlt. If the liposomes were preincubated with PSA for 10-15 min and then Mlt was added, decrease in activity was substantially lower and was evident only at high excess of PSA in the solution. Addition of 2 M NaCl to the outer medium was shown to decrease the activity if the peptide insignificantly (Fig. 4b).
Fig. 4. Time dependence of calcein leakage (% of maximum leakage induced by addition of 10% Triton X-100) from liposomes of different composition under the action of different peptides: DOPC, Mlt, L/P = 85 : 1 (a); DOPC/DOPG (2 : 1), Mlt, L/P = 79 : 1 (b); DOPC, latarcin Ltc1K, L/P = 32 : 1 (c); DOPC/DOPG (2 : 1), latarcin Ltc1K, L/P = 23 : 1 (d). Curves of dye leakage are given for the absence of PSA (1), for the case when a complex between peptide and PSA was formed beforehand (2), in the presence of PSA in the medium (3), and in the presence of 2 M NaCl (4).
Membranolytic activity of Ltc1K. Compared to Mlt, Ltc1K exhibited lower membranolytic activity toward liposomes. Kinetics of calcein leakage from liposomes differed for Ltc1K. Even 20 min after addition of the peptide, the curve of fluorescence intensity of calcein did not reach a plateau. Therefore, in further discussion the fluorescence level (extent of leakage of the dye) observed at 10 min after addition of the latarcin is taken into consideration (Fig. 4b). Latarcin Ltc1K destroyed 30-50% of DOPC liposomes at L/P ratio 32 : 1. For preformed Ltc1K/PSA complex, a substantially higher decrease in membranolytic activity was observed compared to the experiment where the peptide was added to the liposomes in the medium containing PSA (Fig. 4c). This effect is comparable to that observed for Mlt except the initial level of activity of Ltc1K was lower. Irrespective of its high positive charge, this peptide only weakly perturbed DOPC/DOPG liposomes (Fig. 4d). This is in accordance with data obtained previously for liposomes formed of DOPE/DOPG mixture [11].
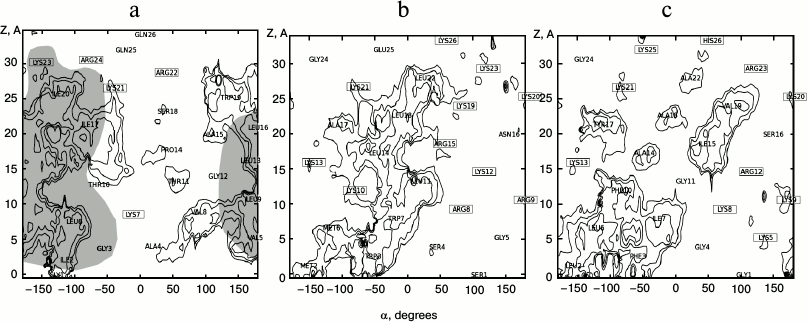
Hydrophobic–hydrophilic properties of the molecular surface of the peptides. Two-dimensional maps of the distribution of molecular hydrophobicity potential (MHP) over the surface of α-helical segments of the peptides were constructed to explain the observed effects (Fig. 5). According to the literature [15, 31], Mlt forms α-helix with a flexible segment incorporating the Gly12-Pro14 residues. The most hydrophobic regions of the peptide are formed by Ile2, Val5, Leu6, Val8, and Leu9 residues at the N-terminus (Gly1-Thr11) and Leu16, Ile17, and Ile20 at the C-terminus (Pro14-Gln26). Exactly these fragments (colored in gray in Fig. 5a) participate in tetramer formation. It is of note that in the contact area there is no charged residue.
Fig. 5. Two-dimensional maps of distribution of hydrophobic regions at the surface of α-helices of Mlt (a), latarcin Ltc1K (b), and latarcin Ltc2a (c). Hydrophobicity is represented by the values of molecular hydrophobicity potential (MHP). Hydrophobic regions (MHP > 0) are indicated with isopotential lines separated by 0.25 MHP units. The angle (α) of rotation around the axis of the helix is indicated on the abscissa. Displacement along the Z-axis is drawn on the ordinate. The locations of amino acid residues are indicated on the map, and the names of charged residues are enclosed. Contacts between monomers of Mlt under conditions of oligomerization are marked by gray filling (based on analysis of X-ray structure of Mlt, 2MLT).
Ltc2a forms a pair of α-helical fragments (according to NMR data [12]) displaying different properties. At the surface of the N-terminal α-helix, a hydrophobic zone formed by the Phe3, Leu6, Ile7, and Phe8 residues is present. In the C-terminal helix there is no distinct division into hydrophobic and hydrophilic zones. Only small islands of hydrophobicity are observed in the vicinity of the Ile15 and Val17 residues. Unlike Mlt, charged residues are distributed evenly over the whole sequence.
Ltc1K forming a single α-helix without kinks (according to NMR data [11]) is characterized by a narrow hydrophobic pattern elongated along the axis of the α-helix. It is constituted by residues Met2, Trp3, Leu11, Leu14, Leu18, and Leu22. It should be mentioned the charged residues Arg8, Lys12, Arg15, Lys19, and Lys23 are located at the border of the hydrophobic zone.
DISCUSSION
In combat against pathogenic microorganisms, antimicrobial peptides are now considered as alternatives to classical antibiotics and as perspective means against cancer as well. Behavior of peptides in model systems is commonly studied. However, their interaction with oligosaccharides and sialic acid in particular is not given adequate attention. Moreover, the reasons are not completely clear as to why similar antimicrobial peptides act selectively against well-defined types of cells.
Taking as an example as many as three LCP, whose hydrophobic properties and structure have much in common, the present work demonstrates the peptides interact drastically differently with PSA and phospholipid liposomes. In our view, these differences can be attributed to individual tendencies of the peptides toward helix formation and distribution of charged residues and hydrophobic–hydrophilic regions over the surface of the formed α-helix. Of the peptides investigated, Mlt exhibits the highest membranolytic activity and the lowest selectivity. Particularly, it turned into helix when tiny amounts of PSA were added, and it effectively destroyed liposomes of any lipid composition. The basic property of this peptide is its pronounced amphiphilicity and presence of a cluster of positive charges, as well as ability to form a tetramer [31]. It is supposed that screening of the charges in this cluster by anions gives rise to helix formation and tetramerization of the peptide. The latter is favored by an increase in the ionic strength of the solution. Polyanions have a larger effect. Effectiveness of association increases in the series: Cl– < SO42– < PO43– [31]. It should be noted that the concentration of negative charges in solution required for substantial tetramerization of Mlt to occur is approximately 300 mM in experiments like this one [31]. When PSA was added, the effective concentration of negative charges was ~1.5 mM, which is substantially lower than that of anions in salts that are required to turn this peptide into a tetramer. A similar effect has been exemplified previously by heparin, heparan sulfate, and dermatan sulfate. All of these induced helix formation in Mlt similarly [25]. It can be supposed that for the case of polyanions a more effective screening of the charged moieties of the peptides is attained due to selective binding. A shift of the maximum position of fluorescence emission of the Trp19 residue shows that, when Mlt interacts with PSA, it turns into oligomers. The extent of helix formation in all three peptides interacting with PSA in PBS was lower than in aqueous solutions (according to CD data), but Mlt did not form oligomers.
It has been noted previously that the behavior of Mlt under conditions of its interactions with PSA differs from its behavior in organic solvents (e.g. in methanol or trifluoroethanol), where the peptide becomes helical but without forming a tetramer [15]. Melittin forms stable complexes with PSA, whose size, according to SAXS data, is somewhat higher than that of the tetramer of Mlt. This indicates that Mlt in the complex is in the tetrameric form and surrounded by several molecules of PSA-7. Our model for the PSA/Mlt complex differs from one suggested earlier for Mlt with heparan sulfate. According to it, the molecules of the peptide in helical conformation were supposed to cover evenly the long oligosaccharide chain, adopting an open conformation [25]. Probably, for oligomerization of Mlt to be seeded a single polyanion or even its part is required. This is reflected in the CD spectra. However, the most effective interaction is attained for ratio close to 1 : 1 for PSA-7. Because the size of the complexes with PSA-7 does not vary when the peptide/PSA ratio is changed, it can be supposed that the interaction of PSA with Mlt is specific.
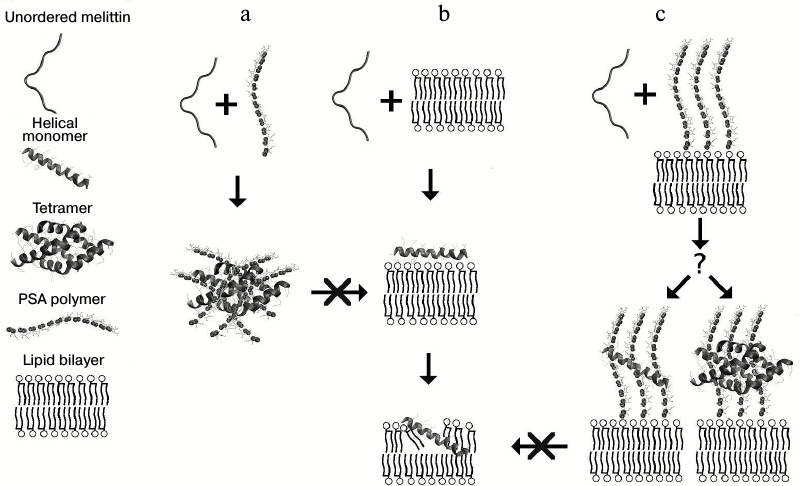
PSA inhibits membranolytic activity of Mlt, but a significant effect is attained when the peptide is preincubated with PSA. This argues in favor of the above-mentioned hypothesis of Mlt being in the tetrameric form in the complexes. When Mlt is added to the liposomes in a medium containing PSA, the peptide can interact with the liposomes, avoiding the stage of complex formation with PSA. It should be noted that activity of Mlt toward liposomes formed of lipid mixture is lower than that to DOPC liposomes. This can be explained by strong attraction of the peptide to charged surface moieties of the lipids, preventing penetration of the peptide into the hydrophobic region of the bilayer. The most important property of Mlt explaining its behavior is exactly its capacity to form a stable tetramer. A schematic diagram of possible routes of interaction of Mlt with PSA and liposomes is presented in Fig. 6.
Concerning Ltc1K, there is still no definite data supporting the possibility of its oligomerization. However, it is supposed that this peptide can form ion-conducting pores in a membrane [11]. Of note, the α-helix formed by Ltc1K possesses noticeable amphiphilicity compared to the two other peptides (Fig. 6b). The charged residues are located evenly on its hydrophilic part. The behavior of the peptide on interaction with liposomes depends upon their composition. PSA induces prominent helix formation in the peptide, contrastingly to Ltc2a. One can say that PSA decreases its membranolytic activity, as observed for Mlt. However, a decrease by a factor of 2-3 was observed for Mlt, while for Ltc1K it was 1.5-2-fold. It is supposed that this peptide interacts with PSA weaker. Possible reasons for this are: 1) absence of a local charge cluster; 2) inability to form a tetramer, in contrast to Mlt; 3) clear amphiphilicity, shifting the equilibrium toward interaction with liposomes even in the presence of PSA.
Fig. 6. Schematic diagram of interaction of Mlt with PSA and liposomes. Formation of inactive tetramer of Mlt in the presence of PSA (a), interaction of the peptide with lipid bilayer (b), and hypothetical mechanism of protection of cells from LCP by anionic polysaccharides (c).
Notwithstanding the similarity to Mlt in hydrophobic properties of the α-helix formed in the presence of phospholipid membranes and micelles, Ltc2a did not show any conformational reorganization when PSA was added to the solution. Detailed analysis of the distribution of MHP values over the surface of the Ltc2a peptide revealed a number of significant differences from Mlt: 1) this peptide features a smaller hydrophobic region; 2) unlike Ltc1K and Mlt, this region is not elongated along the axis of the helix, making impossible formation of a tetramer of Mlt type and strongly reducing the tendency toward helix formation compared to Ltc1K.
Thus, it is demonstrated that three peptides belonging to the same class can interact significantly differently with liposomes of different composition and with PSA. Taking Mlt as an example, it has been shown that anionic polymers are capable of modulating significantly the activity of the LCP. Taking into account that a variety of oligo- and polysaccharides is present on the surface of most biological membranes, it can be assumed that one of their functions is protection of cells from attack of different membranolytic agents, including peptides. A peptide–polymer interaction at the cell surface can be specific. This allows selectively and effectively protecting a cell from various threats. In turn, diversity of LCP can be also be related to the variety of their targets. The behavior of a peptide is determined by not only the extent of its amphiphilicity and net positive charge, but also the definite amino acid sequence, conformational lability, and distribution of physicochemical properties over the surface of the α-helix.
The authors thank the workers of the Laboratory of Oxylipins of the Institute of Bioorganic Chemistry of the Russian Academy of Sciences for samples of Mlt and PSA, A. A. Vassilevski for providing samples of latarcins, V. V. Volkov together with the staff of the Laboratory of SAXS of the Institute of Crystallography for conducting SAXS experiments, and I. A. Boldyrev for help in organizing the experiments with liposomes. Especially, the authors are thankful to A. S. Arseniev, V. V. Bezuglov, and N. M. Gretskaya for attention to the work and discussion of the results. P. Dubovskii thanks N. Kuznetsova for her critical comments on the English translation of the article. R. G. Efremov expresses his gratitude to the scientific foundation “Scientific Investigation of University High Economics School” for support.
This work was fulfilled under support of the program of the Presidium of the Russian Academy of Sciences “Molecular and Cell Biology” and the Russian Foundation for Basic Research.
REFERENCES
1.Corzo, G., Bernard, C., Clement, H., Villegas, E.,
Bosmans, F., Tytgat, J., Possani, L. D., Darbon, H., and Alagon, A.
(2009) Biochim. Biophys. Acta, 1794, 1190-1196.
2.Dubovskii, P. V., Vassilevski, A. A., Samsonova, O.
V., Egorova, N. S., Kozlov, S. A., Feofanov, A. V., Arseniev, A. S.,
and Grishin, E. V. (2011) FEBS J., 278, 4382-4393.
3.Feofanov, A. V., Sharonov, G. V., Dubinnyi, M. A.,
Astapova, M. V., Kudelina, I. A., Dubovskii, P. V., Rodionov, D. I.,
Utkin, Y. N., and Arseniev, A. S. (2004) Biochemistry (Moscow),
69, 1148-1157.
4.Hoskin, D. W., and Ramamoorthy, A. (2008)
Biochim. Biophys. Acta, 1778, 357-375.
5.Vassilevski, A. A., Kozlov, S. A., and Grishin, E.
V. (2009) Biochemistry (Moscow), 74, 1505-1534.
6.Kini, R. M., and Doley, R. (2010) Toxicon,
56, 855-867.
7.Konshina, A. G., Dubovskii, P. V., and Efremov, R.
G. (2012) Curr. Protein Pept. Sci., 13, 570-584.
8.Dubovskii, P. V., Konshina, A. G., and Efremov, R.
G. (2014) Curr. Med. Chem., 21, 270-287.
9.Dubovskii, P. V., Li, H., Takahashi, S., Arseniev,
A. S., and Akasaka, K. (2000) Protein Sci., 9,
786-798.
10.Dubovskii, P. V. (2012) Eur. Biophys. J.,
41, 1077-1084.
11.Dubovskii, P. V., Volynsky, P. E., Polyansky, A.
A., Karpunin, D. V., Chupin, V. V., Efremov, R. G., and Arseniev, A. S.
(2008) Biochemistry, 47, 3525-3533.
12.Dubovskii, P. V., Volynsky, P. E., Polyansky, A.
A., Chupin, V. V., Efremov, R. G., and Arseniev, A. S. (2006)
Biochemistry, 45, 10759-10767.
13.Kozlov, S. A., Vassilevski, A. A., Feofanov, A.
V., Surovoy, A. Y., Karpunin, D. V., and Grishin, E. V. (2006) J.
Biol. Chem., 281, 20983-20992.
14.Vassilevski, A. A., Kozlov, S. A., Zhmak, M. N.,
Kudelina, I. A., Dubovskii, P. V., Shatursky, O. Y., Arseniev, A. S.,
and Grishin, E. V. (2007) Russ. J. Bioorg. Chem., 33,
376-382.
15.Raghuraman, H., and Chattopadhyay, A. (2007)
Biosci. Rep., 27, 189-223.
16.Eisenberg, D., Wilcox, W., and McLachlan, A. D.
(1986) J. Cell. Biochem., 31, 11-17.
17.Vorontsova, O. V., Egorova, N. S., Arseniev, A.
S., and Feofanov, A. V. (2011) Biochimie, 93,
227-241.
18.Gajski, G., and Garaj-Vrhovac, V. (2013)
Environ. Toxicol. Pharmacol., 36, 697-705.
19.Bhunia, A., Domadia, P. N., Torres, J., Hallock,
K. J., Ramamoorthy, A., and Bhattacharjya, S. (2010) J. Biol.
Chem., 285, 3883-3895.
20.Domadia, P. N., Bhunia, A., Ramamoorthy, A., and
Bhattacharjya, S. (2010) J. Am. Chem. Soc., 132,
18417-18428.
21.Bhunia, A., Saravanan, R., Mohanram, H., Mangoni,
M. L., and Bhattacharjya, S. (2011) J. Biol. Chem., 286,
24394-24406.
22.Traving, C., and Schauer, R. (1998)
CMLS, Cell. Mol. Life Sci., 54, 1330-1349.
23.Janas, T., and Janas, T. (2011) Biochim.
Biophys. Acta - Biomembranes, 1808, 2923-2932.
24.Epand, R. M., Epand, R. F., Arnusch, C. J.,
Papahadjopoulos-Sternberg, B., Wang, G., and Shai, Y. (2010)
Biochim. Biophys. Acta, 1798, 1272-1280.
25.Klocek, G., and Seelig, J. (2008)
Biochemistry, 47, 2841-2849.
26.Johnson, W. C. (1999) Proteins:
Structure, Function, and Bioinformatics, 35,
307-312.
27.Efremov, R. G., Gulyaev, D. I., Vergoten, G., and
Modyanov, N. N. (1992) J. Protein Chem., 11, 665-675.
28.Pyrkov, T. V., Chugunov, A. O., Krylov, N. A.,
Nolde, D. E., and Efremov, R. G. (2009) Bioinformatics,
25, 1201-1202.
29.Ghose, A. K., Viswanadhan, V. N., and Wendoloski,
J. J. (1998) J. Phys. Chem. A, 102, 3762-3772.
30.Perez-Paya, E., Houghten, R. A., and Blondelle,
S. E. (1995) J. Biol. Chem., 270, 1048-1056.
31.Tatham, A. S., Hider, R. C., and Drake, A. F.
(1983) Biochem. J., 211, 683-686.