Properties of Hybrid Complexes Composed of Photosynthetic Reaction Centers from the Purple Bacterium Rhodobacter sphaeroides and Quantum Dots in Lecithin Liposomes
V. E. Zagidullin1, E. P. Lukashev1*, P. P. Knox1, N. Kh. Seifullina1, O. S. Sokolova1, E. V. Pechnikova2, H. Lokstein3, and V. Z. Paschenko1
1Biology Faculty, Lomonosov Moscow State University, 119991 Moscow, Russia; fax: (495) 939-1115; E-mail: lukashev@biophys.msu.ru2Shubnikov Institute of Crystallography, Russian Academy of Sciences, Leninsky pr. 59, 119333 Moscow, Russia; E-mail: eugenia.pechnikova@gmail.com
3Institute of Molecular, Cell and Systems Biology, Glasgow Biomedical Research Centre, 120 University Place, University of Glasgow, G12 8TA Glasgow, UK; E-mail: Heiko.Lokstein@glasgow.ac.uk
* To whom correspondence should be addressed.
Received May 22, 2014; Revision received June 23, 2014
Quantum dots (QDs) can absorb ultraviolet and long-wavelength light energy much more efficiently than natural light-harvesting proteins and transfer the excitation energy to photosynthetic reaction centers (RCs). Inclusion into liposomes of RC membrane pigment–protein complexes combined with QDs as antennae opens new opportunities for using such hybrid systems as a basis for artificial energy-transforming devices that potentially can operate with greater efficiency and stability than devices based only on biological components. RCs from Rhodobacter sphaeroides and QDs with fluorescence maximum at 530 nm (CdSe/ZnS with hydrophilic covering) were embedded in lecithin liposomes by extrusion of a solution of multilayer lipid vesicles through a polycarbonate membrane or by dialysis of lipids and proteins dispersed with excess detergent. The dimensions of the resulting hybrid systems were evaluated using dynamic light scattering and by transmission cryoelectron microscopy. The efficiency of RC and QD interaction within the liposomes was estimated using fluorescence excitation spectra of the photoactive bacteriochlorophyll of the RCs and by measuring the fluorescence decay kinetics of the QDs. The functional activity of the RCs in hybrid complexes was fully maintained, and their stability was even increased.
KEY WORDS: purple bacteria, photosynthetic reaction center, quantum dots, liposomes, energy transferDOI: 10.1134/S0006297914110054
Abbreviations: BChl, bacteriochlorophyll; BPheo, bacteriopheophytin; D-D, detergent- and dialysis-based method of liposomes preparation; RC, photosynthetic reaction center; QDs, quantum dots.
A photosynthetic reaction center (RC) is a natural mesoscopic structure
with dimensions of ~10 nm [1-3]. The main cofactors involved in the light energy
conversion into electrochemical potential in pigment–protein
complexes of RCs are porphyrins: (bacterio)chlorophylls, photoactive
dimers of (bacterio)chlorophylls, (bacterio)pheophytin, and some other
auxiliary pigments. The structure of cofactors optimized during
billions of years of evolution as well as their position in the protein
allows photosynthetic RCs to convert light energy into electrochemical
potential with an efficiency of ~100%. According to calculations, the
efficiency coefficient of hybrid phototransformers based on native RCs
is significantly (several times) higher than the efficiency coefficient
of existent silicon solar cells, for which the limiting values are
12-15% [4].
However, for creating a RC-based hybrid energy transformer it is necessary to solve some problems. First, it is necessary to obtain a high stability of photosynthetic pigment–protein complexes in vitro, because without photoprotection and repair systems they are subject to pronounced photodestruction. Second, the solar light absorption by the pigment–protein complexes of RCs is not equally efficient throughout the whole spectrum: the main absorption bands of the RC pigments are in the blue and near infrared regions of the spectrum, and it is important to fill the optical transparency window in the ultraviolet and visible range. Light harvesting can be increased, in particular, by creating a hybrid structure composed of RCs and semiconductor nanocrystals with very high extinction just in the range of UV and visible absorption regions.
Fluorescent semiconductor nanocrystals – quantum dots (QDs) – have unique optical characteristics such as a large absorption section, extremely high photostability, wide absorption spectrum, and a narrow band of photoluminescence. The quantum yield of QD fluorescence reaches 70%, and their extinction coefficient is tenfold higher than the extinction coefficients of organic dyes [5, 6]. Moreover, the emission maximum of QDs is determined by their diameter. Particles of cadmium selenide coated with a zinc sulfide envelope (CdSe/ZnS nanocrystals) with diameters of the nucleus (CdSe) from 2.5 to 6 nm can emit fluorescence quanta in the range of 480-600 nm [6, 7]. Coating QDs with an additional envelope of bi- or trifunctional polymers makes them water-soluble due to polar groups and allows them to bind to biomolecules [8, 9]. These properties of QDs underlie their growing application in biology and medicine [6, 10].
Recent studies have shown that QDs can be used as additional light-harvesters for natural pigment–protein complexes, including photosynthetic RCs of bacteria, and can ensure high (up to 90%) efficiency of energy transfer. Increasing fluorescence of acceptors (photosynthetic proteins) can reach 4-5 units [11, 12]. We have shown that using QDs makes it possible to increase the efficiency of light absorption of synthetic porphyrins by Förster’s inductive-resonance mechanism of energy transfer [13]. This allows us to consider the creation of hybrid energy phototransformers with increased efficiency as a technically solvable problem.
The stability (regarding time, temperature, etc.) of such hybrid systems can be increased by embedding them into liposomes. Liposomes are known to model the natural environment of membrane proteins and have long been successfully used for this purpose [14]. Because of interaction with environmental lipids, the transmembrane helices of membrane proteins are sensitive to lipid characteristics such as lateral wrapping, thickness of the hydrophobic area, and charge of the heads. This can influence the structure and functions of intact membrane proteins, in particular through lateral association of transmembrane segments and their inclination to the normal to the bilayer [15]. The lipid bilayer protects membrane proteins against denaturation and provides the environment required for interaction between different enzymes involved in catalytic cycles. In many cases, lipids also regulate catalytic activities of the enzymes [16, 17]. The interaction of the membrane protein and the environmental lipids can be a prerequisite for providing a conformational mobility of the enzyme necessary for its functioning [18]. All this is also true for photosynthetic pigment–protein complexes, including purple bacterial RCs responsible for the primary separation of electric charges. The lipid composition of photosynthetic membranes of bacteria varies depending on conditions of their growth, in particular, on oxygen content, but phosphatidylcholine is always one of major components of the lipids [19, 20]. Some lipids remain bound to the proteins of RCs isolated from the membrane even after intensive purification. The RCs from Rhodobacter (Rb.) sphaeroides retain tight binding with three lipid molecules: cardiolipin, phosphatidylcholine, and glucosyl galactosyl diacylglycerol [21]. This suggests that specific interactions exist between phospholipids and RCs. Thermodynamic and kinetic parameters of electron transfer via the quinone acceptor of the purple bacteria RCs were found to depend on the presence of physiologically important lipids (phosphatidylcholine, phosphatidylglycerol, phosphatidylethanolamine, cardiolipin) in their environment [22, 23]. The mechanism of the lipid influence on the photosynthetic energy transformation is still unclear, although it was supposed [22] that anionic phospholipids should influence the primary quinone acceptor of RCs with the appearance of a long-living charge separated state. However, earlier we showed that electron transfers in RCs, in particular processes involving quinone receptors, are essentially associated with the structural and dynamic state of the RCs. Studies on temperature and hydration degree dependences revealed clear correlations between the intramolecular mobilities of different parts of photosynthetic preparations and changes in the efficiency of electron transfer in the photosynthetic chain of electron transfer [24-26].
The purpose of the present work was to embed RCs and nanocrystals capable of absorbing light over a wide spectral range into liposomes prepared from phosphatidylcholine (lecithin) and to study the energetic interaction of components of such hybrid complexes within the liposome structure.
MATERIALS AND METHODS
Cells of the purple bacterium Rb. sphaeroides were broken with an ultrasonic disintegrator. Chromatophores separated by centrifugation were incubated for 30 min at 4°C in 0.01 M sodium phosphate buffer (pH 7.0) supplemented with 0.5% of the zwitterionic detergent lauryl dimethylamine oxide (LDAO). Then the chromatophores were centrifuged at 144,000g for 90 min at 4°C. The fraction of RCs that was present in the supernatant was separated by chromatography on a column with hydroxyapatite as described in detail previously [27]. The concentration of the resulting RCs suspended in 0.01 M sodium phosphate buffer (pH 7.0) supplemented with 0.05% LDAO was ~10 µM. Photoreactions were studied using a single-beam differential spectrophotometer (in the Qy absorption band of bacteriochlorophyll (BChl) P870 (at 870 nm).
CdSe/ZnS quantum dots with a luminescence maximum at ~530 nm functionalized with a hydrophilic polymeric coating with carboxyl COOH-groups were used (Rusnanotech, Russia).
To prepare liposomes, L-α-lecithin from soybean (Sigma, USA) was used. Two methods were used. The first was a comparatively new method of preparing single-walled vesicles by repeated pressing (extrusion) of a multilayer lipid vesicle suspension through a porous polycarbonate membrane (in our case the pore size was 0.1 µm) in a special device – Extruder (Avanti Polar Lipids, Inc., USA). In another method, both detergent and dialysis were used (D-D).
The method of preparing vesicles using detergent and dialysis (D-D) included the following steps: 1) 50 mg lecithin was dissolved in 0.5 ml of chloroform and evaporated; 2) the lipid film was covered with 2 ml of 50 mM Tris-HCl buffer (pH 8) containing 2% sodium cholate; 3) the lipid suspension was sonicated with a UZDN-2T ultrasonic disintegrator (Soyuz-Pribor, Russia) until it became clear; 4) a 0.5-ml aliquot of the resulting lipid solution in Tris buffer was supplemented with 0.2 ml of Tris buffer, sonicated for 5 s, and used for preparing lecithin liposomes; 5) a 0.5-ml aliquot of the resulting lipid solution in Tris buffer was supplemented with 0.1 ml of Tris buffer and 0.1 ml of RCs (42 µM), sonicated for 5 s, and used for preparing RC-containing proteoliposomes; 6) a 0.5-ml aliquot of the resulting lipid solution in Tris buffer was supplemented with 0.1 ml of Tris buffer and 0.1 ml of quantum dots (43 µM), sonicated for 5 s, and used for preparing QD-containing liposomes; 7) a 0.5-ml aliquot of the resulting lipid solution in Tris buffer was supplemented with 0.1 ml of RCs (42 µM) and 0.1 ml of quantum dots (43 µM), sonicated for 5 s, and used for preparing RC/QD-containing proteoliposomes; 8) the samples (4-7) were placed into dialysis bags and on a stirrer placed into a refrigerator to dialyze against 1 liter of 50 mM Tris-HCl buffer (pH 8).
Using the extruder, 50 mg lecithin was also dissolved in 0.5 ml of chloroform (analytical grade) and the solvent was slowly evaporated. Then 2 ml of 50 mM Tris-HCl buffer (pH 8) was added and the lipid was dissolved in the buffer using a vortex mixer. To accelerate dissolution, the suspension was frozen and thawed repeatedly. Solutions containing lipids and QDs, lipids and RCs, and also lipids and mixtures of QDs with RCs were prepared in the same ratios as described above for the D-D method, but instead of the ultrasonic treatment the solutions were mixed with a vortex mixer. Later, the lipid solution or lipids containing QDs, RCs, and RCs/QDs were thirteen times pressed through a membrane with 0.1 µm pores using the extruder.
The mean size of the resulting liposomes was measured with a Z-sizer (Malvern, USA) that allowed us to determine the hydrodynamic radius of the particles by changes in their light scattering. Each measurement was repeated five times, and the results were averaged.
Samples for transmission cryoelectron microscopy were prepared in the Shubnikov Institute of Crystallography, Russian Academy of Sciences, using a Vitrobot Mark IV device (FEI, Netherlands) that automatically controls the humidity and temperature. Technical details of the device and method are described in [28, 29]. A support (Quantifoil, Germany) pretreated for 45 s in a smoldering charge atmosphere was placed into a chamber with controlled temperature and ultrasound emitter to create humidity (up to 100% relative humidity). On the support, 3 µl of the sample suspension was placed. The excess fluid was automatically removed with paper filters. Thin films produced in the support holes were instantly frozen by quick submerging of the sample into liquid ethane. In our experiments, the samples were pretreated at 37°C and 100% relative humidity.
The supports with the samples were examined with a CM-12 transmission electron microscope (Philips, Netherlands). During the experiment the voltage was 120 kV and the pressure <0.2·10–3 Pa. Microphotographs were obtained using the technology of imaging plates under standard conditions of exposure in the regime of “low dose” of electrons per Å2.
For electrophoresis, 10-µl aliquots of freshly prepared aqueous samples (for the doubling experiment a twofold greater volume was taken) were placed into 1% agarose gel prepared in 0.25× TBE buffer (54 g Tris, 27.5 g boric acid, 20 ml of 0.5 M EDTA, pH 8.0) and then placed into an electrophoretic cell with 0.5× TBA buffer. The gel was kept for 60 min at a voltage of 80 V using a BioRad electrophoretic system (USA) and then photographed under ultraviolet illumination on a GeneGenius system (USA).
The fluorescence excitation spectra of the samples were recorded using a SPEX Fluorolog II spectrofluorimeter (Jobin Yvon, France). Fluorescence kinetics were measured using a pulse fluorimeter excited with picosecond light pulses (λex = 532 nm, FWHM ~ 20 ps) at a frequency of 1 Hz from a PIKAR-1 laser (Moscow State University, Russia). A sample of 0.5 ml was placed into a cuvette of 2 mm thickness, the optical density at λ = 532 nm being 0.1. The fluorescence kinetics were recorded using an AGAT SFZ electro-optical chamber (Proton, Russia) connected with a computer through a C7041 multichannel matrix detector and a C7557 controller (HAMAMATSU, Japan). The time resolution of the system was ~2 ps. During the experiment, the samples were stirred with a magnetic stirrer. To improve the signal/noise ratios, 50 kinetic traces were accumulated and averaged. The accumulated experimental kinetics were approximated bi-exponentially by modeling the curve as I(t) = A1exp(–t/τ1) + A2exp(–t/τ2) and the apparent characteristic function of the fluorimeter, where I(t) is the fluorescence change of the sample with time; and τ1, τ2 and A1, A2 are lifetimes and amplitudes of the fast (1) and slow (2) components of the fluorescence decay kinetics (A1 + A2 = 1).
RESULTS AND DISCUSSION
The main advantages of preparing liposomes by extrusion through a porous polycarbonate membrane are the simplicity of the method and the possibility to widely vary the size of vesicles by changing the size of the polycarbonate filter pores (from tens of nanometers to microns). Our measurements by dynamic light scattering of the size of lecithin liposomes prepared by extrusion through the porous polycarbonate membrane with pores of 0.1 µm showed that the average size of the particles was ~100 nm. However, the extrusion of the lipid mixture containing the RC proteins resulted in insufficiently homogenous proteoliposomes, which indicated poor incorporation of the proteins into the structure of the vesicles. For preparing proteoliposomes, it was better to use the other method with the lipid and the protein dispersed in excess detergent and subsequent removal of the detergent, e.g. by dialysis (the detergent-dialysis method, D-D). The lipid bilayer thickness was in good correlation with the size of the membrane proteins that promoted the formation of stable closed structures, and the orientation degree of the proteins on such incorporation was >80%. The proteoliposome diameter in this case depended on the nature of the detergent and on the rate of its removal. We used this approach for preparation of proteoliposomes containing proteins of the RCs from the purple bacterium Rb. sphaeroides and quantum dots. The average diameter of lecithin liposomes prepared by the D-D method was ~40 nm. This value correlates with results of other studies. In particular, it was shown in [30] that the size of proteoliposomes depended on the lipid/protein ratio in the initial mixture: the greater the amount of lipid, the smaller the liposomes. For the lipid dioleoylphosphatidylcholine and RCs from Rb. sphaeroides at ratios of 8000 : 1, 2000 : 1, and 1000 : 1, the hydrodynamic radii obtained in this work for the particles were, respectively, 21 ± 2, 35 ± 3, and 55 ± 6 nm. In our case the average hydrodynamic radius of the RC-containing liposomes was 39 ± 4 nm, of the liposomes containing RC and QD – 38 ± 3 nm, and of the liposomes containing only QD – 27 ± 2 nm.
The orientation of RCs from Rb. sphaeroides in proteoliposomes obtained by dialysis of the detergent sodium cholate was studied in work [31] by the functional interaction of RCs with exogenously added cytochrome c, which is a natural donor of electrons for the photoactive BChl. The transmembrane orientation of the RCs within the proteoliposomes showed, that the donor part of the RC (the BChl dimer – P870) exposed to the outside was increased from 45 to 85% with increase in the size of the vesicles.
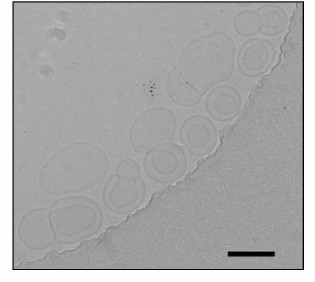
We studied the liposomes prepared by the two methods by electron microscopy. The liposomes prepared by extrusion were characterized by two clearly pronounced layers, and the inner layer diameter was ~60 nm and that of the outer layer was ~120 nm (Fig. 1). Because of the large size of such liposomes, the image is only an optical section, and the greater part of their surface is invisible, as well as the QDs that occur outside of the focal plane, which explains their small number in the photograph.
Fig. 1. Transmission electron microphotograph of liposomes prepared by the extrusion method. The marker corresponds to 120 nm.
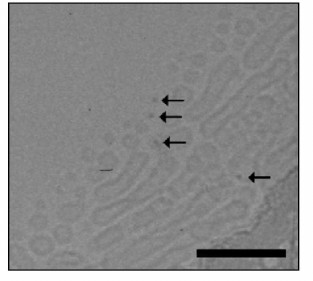
Liposomes prepared by dialysis (Fig. 2) were typically spheres with a diameter of ~25 nm and also larger elongated structures with diameters of 25-30 nm and lengths of 120-250 nm. The latter shape seemed to be caused by fusion of smaller particles on the support. In images of samples with quantum dots added during their preparation denser objects can be seen – the quantum dots indicated by arrows. We concluded this based on the following reasons:
– the sizes of these particles are ~5 nm, which is in good agreement with the theoretical size of the QD nucleus corrected for focusing; the same sizes were recorded in [32];
– the detected objects are in direct contact with the liposomes, which is in agreement with the electrophoretic data regarding QD and liposome aggregation (see further). It is interesting to note that QDs interact with the lipid bilayer of the phosphatidylcholine liposome but are not simply encapsulated in it [33].
Fig. 2. Transmission electron microphotograph of liposomes prepared by the D-D method. The label corresponds to 120 nm. Arrows designate QDs.
The interaction of QDs with liposomes in our preparations was also evident by results of electrophoresis in agarose gels. The QDs (in aqueous solution) did not diffuse in the gel under the influence of the electric field – a bright spot of the UV-excited fluorescence of QDs remained at the origin. This was clearly because of the small charge of the individual particles. However, QD-containing liposomes carrying on their surface many charges of lipid polar heads markedly diffused in the electric field. However, the fluorescent spot was blurred obviously due to the liposome distribution in size and weakened as a result of a partial quenching of the QD fluorescence by phospholipids [33, 34]. Liposomes containing QDs and liposomes containing both QDs and RC proteins have virtually the same electrophoretic mobilities.
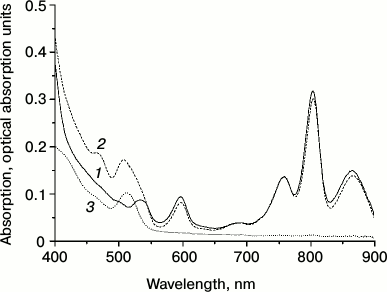
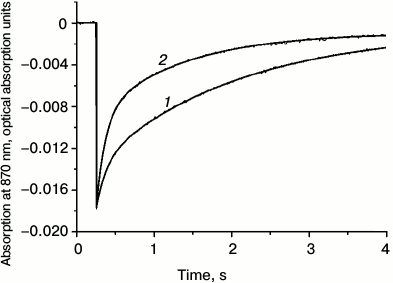
Absorption spectra of RC-containing proteoliposomes, of RC/QD-containing proteoliposomes, and also of quantum dots QD530 with a nucleus of CdSe in aqueous suspension are presented in Fig. 3. The RC absorption spectrum essentially corresponds to the spectrum of the initial RC suspension before the incorporation procedure. However, attention should be given to the disappearance of characteristic absorption bands of carotenoids because of their extraction due to the ultrasonic treatment of the aqueous mixture of RC with lipids in 2% sodium cholate solution that was used by us to improve the dispersion of individual components. If the short-term ultrasonication of the mixture before the dialysis was omitted, the resulting liposomes displayed all absorption bands characteristic for RCs, including the carotenoid band. In vesicles containing only QDs or both QDs and RCs, one can clearly see characteristic absorption bands of the QDs in the region of 400-550 nm. The functional activity of RCs – the ability to reversibly photooxidize the BChl RC (P870) under the pulsed illumination of the proteoliposomes – was also retained (Fig. 4). This was also true for hybrid liposomal structures containing RCs and QDs. The kinetics of photoinduced changes in the P870 absorption presented in Fig. 4 reflect the charge separation between the primary electron donor, P870, and the quinone acceptors (QA, QB) with their subsequent dark recombination. The kinetics recorded in RC-containing vesicular preparations reflecting the dark recombination of P+ and QB– was close to the kinetics for the original RC preparation in aqueous buffer. The recombination kinetics in proteoliposomes containing RCs and QDs was faster than in vesicles containing only RCs without QDs (Fig. 4). Obviously, this is associated with changes in the efficiency of the transient electron stabilization at the quinone acceptor component of the RCs in QD-containing preparations. An effective electrostatic stabilization of electrons at the quinone acceptors QA and QB in RCs of purple bacteria was associated with displacements of protons in their protein environment. Moreover, not only protonated amino acid residues of the nearest environment of quinone cofactors were involved in these processes. It was shown earlier that changes in the charge of the quinones could be accompanied by changes in the pK of protonated amino acids at distances of 15-17 Å [35, 36]. As noted in [37], the rate constant of the dark recombination of the electron from QB– to P+ was sensitive to the ionization state of all charged residues in the environment of QB–. The influence of QDs on the transient stabilization of separated charges seems to indicate that their closeness to RCs in liposomes is sufficient to influence the structural dynamics of peripheral regions of the RCs, which was reflected in the recorded recombination kinetics.
Fig. 3. Absorption spectra of lecithin proteoliposomes with RCs from Rb. sphaeroides (1), proteoliposomes with both RCs and quantum dots (2), and liposomes with only quantum dots (3).
Fig. 4. Kinetics of absorption changes of the photoactive bacteriochlorophyll at 870 nm induced by laser illumination in proteoliposomes containing RCs of Rb. sphaeroides (1) and proteoliposomes containing both RCs and QDs (2).
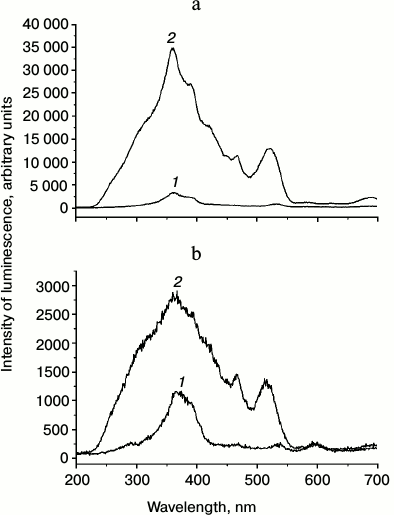
The effective interaction of RCs and QDs was also confirmed by the fluorescence characteristics of the hybrid complexes. Figure 5 (a and b) presents fluorescence excitation spectra of photoactive BChl P870 in proteoliposomes (1) and in the RC/QD-containing liposomal preparations (2) recorded at 775 and 910 nm. The fluorescence excitation spectrum of the RC-containing preparation without QDs virtually coincided with the absorption spectrum of the RCs. The figure also shows a strong increase in the fluorescence signal of the preparation containing both RCs and QDs due to absorption of light energy by quantum dots in the range of 250-550 nm and its efficient transfer to the RCs. Considering that we used QDs with an emission maximum at 530 nm that coincided with the Qx absorption band of bacteriopheophytin (BPheo) in RCs (bacteriochlorophylls in RCs have a maximum of the Qx absorption band at ~590 nm), the fluorescence signal of the hybrid complexes was the highest when luminescence was recorded at 775 nm, i.e. near the BPheo fluorescence maximum (the Qy absorption band of BPheo has a maximum at ~760 nm). The excitation energy trapped by BPheo molecules was transferred to BChl monomers with an absorption maximum in the Qy region of 800 nm and further to the photoactive BChl dimer with an absorption Qy maximum near 870 nm with fluorescence at 910 nm.
Fig. 5. Fluorescence excitation spectra of proteoliposomes containing RCs (1) and in liposomes containing both RCs and QDs (2) recorded at 775 nm (a) and 910 nm (b).
The excited state of QDs was effectively quenched as a result of the interaction of the QDs with the RCs in the liposomes, which was also confirmed by measurements of the QD fluorescence decay kinetics. The fluorescence decay kinetics averaged from 50 measurements were approximated with a biexponential function of the type: A = A1e(–t/τ1) + A2e(–t/τ2), where A1, τ1 are the amplitude and characteristic time, respectively, of the first decay component and A2, τ2 are the amplitude and characteristic time of the second decay component. The average lifetime of the fluorescence decay was characterized by the ratio as follows: Tav = (Α1τ1 + Α2τ2)/(A1 + A2).
Preparations of QD-containing liposomes were supplemented with RCs at concentrations increasing from 1 to 11 µM. Parameters of the obtained kinetic curves (for the biexponential approximation) are given in the table.
Amplitudes (A) and characteristic lifetimes (T) of decay kinetics of the
QD fluorescence in proteoliposomes containing RCs of Rb.
sphaeroides depending on the concentration of RCs added
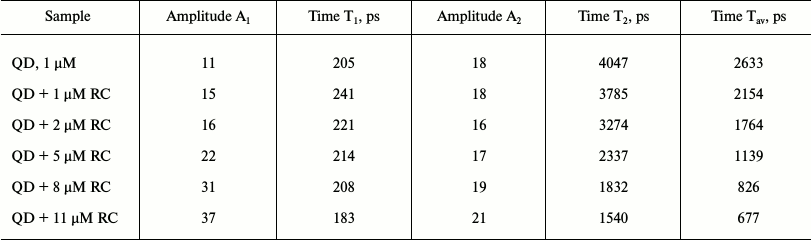
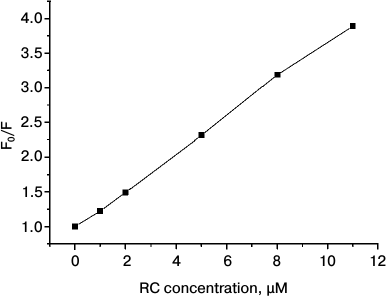
An increase in the amount of RCs was accompanied by a decrease in the characteristic lifetime of the fluorescence decay, which indicated the dynamic character of its quenching. This conclusion was confirmed by the plot of the dependence of fluorescence quenching (ratio of maximal fluorescence to fluorescence observed upon addition of a quencher), i.e. on the RC concentration in Stern–Volmer coordinates (Fig. 6). This plot indicated that this ratio increased with increasing concentration and was linear, which confirms the dynamic character of QD fluorescence quenching [38]. It should be noted that during this experiment QDs were in a liposomal environment (some QDs were encapsulated in the interior of the liposomes, and others were incorporated into the lipid bilayer), whereas RCs were in the aqueous medium. In this case, quenching of QD fluorescence by RCs was lower than during the titration with QDs in solution. This could be expected because QDs localized within liposomes could not interact with the RCs localized outside. The purpose of this experiment was to show that the QD particles incorporated into the lipid bilayer retained their fluorescence properties and could form complexes with RCs localized on the surface of the liposomes. Unfortunately, it was technically impossible to obtain preparations of proteoliposomes with both QDs and RC proteins localized only within the lipid bilayer in different ratios and to perform kinetic measurements. But in liposomal preparations 6 and 7 (see “Materials and Methods”) with the same contents of QDs a noticeable decrease in the lifetime of the QD excited state in the presence of RCs was found (data not presented). This indicates the formation of a molecular QD/RC complex with energy transfer from QDs to RCs within the liposomes.
Fig. 6. Quenching of QD fluorescence in preparations of proteoliposomes with different concentrations of RCs from Rb. sphaeroides in Stern–Volmer coordinates.
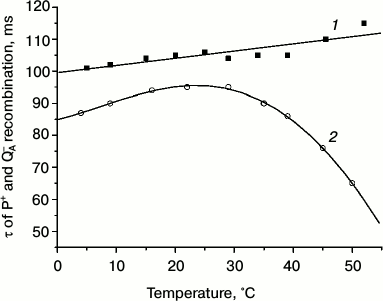
To assess the stability of RC function within proteoliposomes, we studied the temperature dependence of the characteristic lifetimes of dark recombination of P+ and QA– in Rb. sphaeroides RCs at temperatures above room temperature (Fig. 7). This functional parameter is an informative index of the structural-dynamical state of the RCs that determines their activity. We found earlier that in an aqueous suspension of isolated RCs the characteristic time of this reaction decreased with increase in temperature above room temperature (up to 45-50°C the recorded functional effects were reversible [39]), which suggests a decrease in the time of stabilization of the photoseparated charges between P+ and QA– as a result of the increased probability of their recombination. We associate this observation with specific features of the temperature dependence of the state and dynamics of hydrogen bonds influencing the stabilization of separated charges in the environment of the electron transfer cofactors in the RC structure [40, 41].
Fig. 7. Temperature dependence of the characteristic lifetime of dark recombination of photoseparated charges between bacteriochlorophyll P and the primary quinone acceptor for the Rb. sphaeroides RCs within lecithin liposomes (1) and RCs in aqueous buffer (2).
To study the temperature dependence of the P+ and QA– charge recombination in RCs in proteoliposomes, RC preparations were pretreated with o-phenanthroline (to 10–2 M), which inhibits electron transfer from the primary quinone acceptor to the secondary one. The samples were excited by separate light flashes. Figure 7 shows that at room temperature values of the recombination rate for RCs within liposomes and in control RCs in aqueous buffer solution were similar: t1/2 were ~100 and 95 ms, respectively. It should be noted that addition of QDs to preparations of RC-containing proteoliposomes had virtually no influence on the value of this functional parameter. However, during the subsequent heating of RC-containing liposome samples, significant differences were observed from the control (RCs in buffer): the characteristic reaction time not only did not decrease with heating up to 50°C but even slightly increased – to 110 ms (Fig. 7). In other words, the efficiency of charge stabilization in the P+—QA– ion-radical pair did not decrease in RCs in the liposomal environment even on heating up to 50°C, as compared to RCs in aqueous buffer. It seemed that the interaction of RCs with the artificial lipid membrane strongly influenced the temperature dependence of electron transfer within the RC structure. The incorporation of RCs into the lipid bilayer led to a significant increase in its thermostability. This was also directly evidenced by the absence of any sign of the protein degradation in the absorption spectra of proteoliposomes containing Rb. sphaeroides RCs during heating to 50°C. The ratio of the absorption bands of the RC porphyrin pigments in the near IR region (760 nm – BPheo, 800 nm – monomeric BChl, 870 nm – photoactive BChl dimer) was virtually unchanged for RCs in liposomes, whereas at 50°C changes specific for pheophytinization were observed for RCs in aqueous buffer, i.e. BChl molecules lost the central Mg atom, the amplitude of the BPheo absorption bands at 800 and 870 nm decreased, and the amplitude of its absorption band at 760 nm increased.
In the present work the possibility has been shown of creating efficient hybrid constructs composed of phototransforming RCs and inorganic nanocrystals embedded into artificial membranous vesicular particles. Stable structures have been prepared with RCs fully retaining their functional activity and quantum dots effectively transmitting the absorbed energy to the RCs. Reliable functional contact was shown to exist in the QD/RC complex in the lipid bilayer structure, with QDs acting as an additional light-harvesting component. In addition, the hybrid complexes were produced by self-assembly due to electrostatic interactions. The liposomal structure was responsible for the high resistance of their protein component to denaturing exposures that was shown by the influence of temperature on RCs in aqueous buffer and within the artificial membranous system. Note that just the comparative study on functional parameters of the phototransforming activity of RCs in aqueous buffer or within natural or artificial membranous structures is currently an urgent problem for investigations of fundamental regulatory mechanisms of light energy conversion in photosynthetic RCs. On the other hand, it can be stated that liposomal structures possess obvious virtues also for applied developments of light-capturing devices based on natural photosensitive proteins.
This work was supported by the Russian Foundation for Basic Research (projects No. 14-04-01536 and 13-04-00403), by grants from ESF EuroCore, BB/J00823011, as well as from the BMBF, 031A154B (H. L.).
REFERENCES
1.Hu, X., Ritz, T., Damjanovic, A., Autenrieth, F.,
and Schulten, K. (2002) Photosynthetic apparatus of purple bacteria,
Quart. Rev. Biophys., 35, 1-62.
2.Yoder, L. M., Cole, A. G., and Sension, R. J.
(2002) Structure and function in the isolated reaction center complex
of photosystem II: energy and charge transfer dynamics and mechanism,
Photosynth. Res., 72, 147-158.
3.Busch, A., and Hippler, M. (2011) The structure and
function of eukaryotic photosystem I, Biochim. Biophys. Acta,
1807, 864-877.
4.McConnell, I., Li, G., and Brudvig, G. W. (2010)
Energy conversion in natural and artificial photosynthesis, Chem.
Biol., 17, 434-447.
5.Leatherdale, C. A., Woo, W.-K., Mikulec, F. V., and
Bawendi, M. G. (2002) On the absorption cross section of CdSe
nanocrystal quantum dots, J. Phys. Chem. B, 106,
7619-7622.
6.Oleynikov, V. A., Sukhanova, A. V., and Nabiev, I.
R. (2007) Fluorescent semiconductor nanocrystals for biology and
medicine, Ros. Nanotekhnol., 2, 160-173.
7.Micic, O. I., Cheong, H. M., Fu, H., Zunger, A.,
Sprague, J. R., Mascarenhas, A., and Nozik, A. J. (1997) Size-dependent
spectroscopy of InP quantum dots, J. Phys. Chem. B, 101,
4904-4912.
8.Gerion, D., Pinaud, F., Williams, S. C., Parak, W.
J., Zanchet, D., Weiss, S., and Alivisatos, A. P. (2001) Synthesis and
properties of biocompatible water-soluble silica-coated CdSe/ZnS
semiconductor quantum dots, J. Phys. Chem. B, 105,
8861-8871.
9.Pons, T., Medintz, I. L., Sapsford, K. E.,
Higashiya, S., Grimes, A. F., English, D. S., and Mattoussi, H. (2007)
On the quenching of semiconductor quantum dot photoluminescence by
proximal gold nanoparticles, Nano Lett., 7,
3157-3164.
10.Medintz, I. L., and Mattoussi, H. (2009) Quantum
dot-based resonance energy transfer and its growing application in
biology, Phys. Chem. Chem. Phys., 11, 17-45.
11.Nabiev, I., Rakovich, A., Sukhanova, A.,
Lukashev, E., Zagidullin, V., Paschenko, V., Rakovich, Y. P., Donegan,
J. F., Rubin, A. B., and Govorov, A. O. (2010) Fluorescent quantum dots
as artificial antennas for enhanced light harvesting and energy
transfer to photosynthetic reaction centers, Angew. Chem.,
49, 7217-7221.
12.Maksimov, E. G., Lukashev, E. P., Seifullina, N.
Kh., Nizova, G. V., and Paschenko, V. Z. (2013) Photophysical
properties of hybrid complexes consisting of quantum dots and reaction
centers of the purple photosynthetic bacteria Rhodobacter
sphaeroides adsorbed on crystalline mesoporous TiO2
films, Ros. Nanotekhnol., 8, 11-17.
13.Borissevitch, I. E., Parra, G. G., Zagidullin, V.
E., Lukashev, E. P., Knox, P. P., Paschenko, V. Z., and Rubin, A. B.
(2013) Cooperative effects in CdSe/ZnS-PEGOH quantum dot luminescence
quenching by a water soluble porphyrin, J. Luminesc.,
134, 83-87.
14.Gennis, P. (1997) Biomembranes: Molecular
Structure and Functions [Russian translation], Mir, Moscow.
15.Nyholm, T. K. M., Ozdirekcan, S., and Killian, J.
A. (2007) How protein transmembrane segments sense the lipid
environment, Biochemistry, 46, 1457-1465.
16.Sandermannr, H., Jr. (1978) Regulation of
membrane enzymes by lipids, Biochim. Biophys. Acta, 515,
209-237.
17.Latruffe, N., Berrez, J. M., and el Kebbaj, M. S.
(1986) Lipid–protein interactions in biomembranes studied through
the phospholipids specificity of D-beta-hydroxybutyrate dehydrogenase,
Biochimie, 68, 481-491.
18.Lee, S. Y., Lee, A., Chen, J. Y., and MacKinnon,
R. (2005) Structure of the KvAP voltage-dependent K+-channel
and its dependence on the lipid membrane, Proc. Natl. Acad. Sci.
USA, 102, 15441-15446.
19.Onishi, J. C., and Niederman, P. (1982)
Rhodopseudomonas sphaeroides membranes: alterations in
phospholipid composition in aerobically and phototrophically grown
cells, J. Bacteriol., 149, 831-839.
20.Benning, C. (2004) Membrane lipids in anoxygenic
photosynthetic bacteria, Adv. Photosynth. Respir., 6,
83-101.
21.Camara-Artigas, A., Brune, D., and Allen, J. P.
(2002) Interactions between lipids and bacterial reaction centers
determined by crystallography, Proc. Natl. Acad. Sci. USA,
99, 11055-11060.
22.Agostiano, A., Milano, F., and Trotta, M. (2005)
Trapping of a long-living charge separated state of photosynthetic
reaction centers in proteoliposomes of negatively charged
phospholipids, Photosynth. Res., 83, 53-61.
23.Milano, F., Dorogi, M., Szebenyi, K., Nagy, L.,
Maroti, P., Varo, G., Giotta, L., Agostiano, A., and Trotta, M. (2007)
Enthalpy/entropy driven activation of the first interquinone electron
transfer in bacterial photosynthetic reaction centers embedded in
vesicles of physiologically important phospholipids,
Bioelectrochemistry, 70, 18-22.
24.Berg, A. I., Knox, P. P., Kononenko, A. A.,
Frolov, E. N., Khrymova, I. N., Rubin, A. B., Likhtenstein, G. I.,
Goldansky, V. I., Parak, F., Bukl, M., and Messbauer, P. (1979)
Conformational regulation of functional activity in photosynthetic
membranes of purple bacteria, Mol. Biol., 13, 81-89.
25.Kotelnikov, A. I., Likhtenstein, G. I., Fogel, V.
R., Kochetkov, V. V., Knox, P. P., Kononenko, A. A., Grishanova, N. P.,
and Rubin, A. B. (1983) Intramolecular dynamics and electron transfer
in photosynthetic reaction centers. The study by luminescence method,
Mol. Biol., 17, 846-855.
26.Kononenko, A. A., Knox, P. P., Chamorovsky, S.
K., Rubin, A. B., Likhtenstein, G. I., Krupyansky, Y. F., Suzdalev, I.
P., and Goldansky, V. I. (1986) Electron transfer and intramolecular
dynamics of photosynthetic reaction centers, Khim. Fiz.,
5, 795-804.
27.Zakharova, N. I., and Churbanova, I. Y. (2000)
Methods for preparation of reaction centers of photosynthesizing purple
bacteria, Biochemistry (Moscow), 65, 149-159.
28.Bellare, J. R., Davis, H. T., Scriven, L. E., and
Talmon, Y. (1988) Controlled environment vitrification system: an
improved sample preparation technique, J. Electron Microsc.
Tech., 10, 87-111.
29.Frederik, P. M., Stuart, M. C., Bomans, P. H.,
Busing, W. M., Burger, K. N., and Verkleij, A. J. (1991) Perspective
and limitations of cryo-electron microscopy. From model systems to
biological specimens, J. Microsc., 161, 253-262.
30.Milano, F., Italiano, F., Trotta, M., and
Agostiano, A. (2009) Characterization of RC–proteoliposomes at
different RC/lipid ratios, Photosynth. Res., 100,
107-112.
31.Iba, K., Takamiya, K., Arata, H., Toh, Y., and
Nishimura, M. (1984) Transmembrane orientation of reaction centers in
proteoliposomes from Rhodopseudomonas sphaeroides, J.
Biochem., 96, 1823-1830.
32.Shan, G.-Y., Li, D., Feng, L.-Y., Kong, X.-G.,
Liu, Y.-C., Bai, Y.-B., Li, T.-J., and Sun, J.-Z. (2005) Encapsulation
of CdSe/ZnSe quantum dots by liposome complexes, Chin. J. Chem.,
23, 1688-1692.
33.Al-Jamal, W. T., Al-Jamal, K. T., Bomans, P. H.,
Frederik, P. M., and Kostarelos, K. (2008)
Functionalized-quantum-dot-liposome hybrids as multimodal nanoparticles
for cancer, Small, 4, 1406-1415.
34.Generalov, R., Kavaliauskiene, S., Westrom, S.,
Chen, W., Kristensen, S., and Juzenas, P. (2011) Entrapment in
phospholipid vesicles quenches photoactivity of quantum dots, Int.
J. Nanomed., 6, 1875-1888.
35.Lancaster, C. R. D., Michel, H., Honig, B., and
Gunner, M. R. (1996) Calculated coupling of electron and proton
transfer in the photosynthetic reaction center of Rhodopseudomonas
viridis, Biophys. J., 70, 2469-2492.
36.Miksovska, J., Maroti, P., Tandori, J., Schiffer,
M., Hanson, D. K., and Sebban, P. (1996) Distant electrostatic
interactions modulate the free energy level of
QA– in the photosynthetic reaction center,
Biochemistry, 35, 15411-15417.
37.Paddock, M. L., Rongey, S. H., McPherson, P. H.,
Juth, A., Feher, G., and Okamura, M. Y. (1994) Pathway of proton
transfer in bacterial reaction centers: role of aspartate-L213 in
proton transfers associated with reduction of quinone to
dihydroquinone, Biochemistry, 33, 134-145.
38.Lakowicz, J. R. (1999) Principles of
Fluorescence Spectroscopy, Kluwer.
39.Krasilnikov, P. M., Knox, P. P., Lukashev, E. P.,
Paschenko, V. Z., Churbanova, I. Y., Shaitan, K. V., and Rubin, A. B.
(2000) Acceleration of the reaction of photooxidized
bacteriochlorophyll and of reduced primary quinone in reaction centers
of Rb. sphaeroides at T > 300 K, Dokl. Akad. Nauk,
375, 828-830.
40.Krasilnikov, P. M., Mamonov, P. A., Knox, P. P.,
Paschenko, V. Z., and Rubin, A. B. (2007) The influence of hydrogen
bonds on electron transfer rate in photosynthetic RCs, Biochim.
Biophys. Acta, 1767, 541-549.
41.Krasilnikov, P. M., Knox, P. P., and Rubin, A. B.
(2009) Relaxation mechanism of molecular systems containing hydrogen
bonds and free energy temperature dependence of reaction of charges
recombination within Rhodobacter sphaeroides RC, Photochem.
Photobiol. Sci., 8, 181-195.