Highly Specific Hybrid Protein DARPin-mCherry for Fluorescent Visualization of Cells Overexpressing Tumor Marker HER2/neu
K. E. Mironova1,2*, O. N. Chernykh1,3, A. V. Ryabova4, O. A. Stremovskiy1, G. M. Proshkina1, and S. M. Deyev1,2
1Shemyakin–Ovchinnikov Institute of Bioorganic Chemistry, Russian Academy of Sciences, ul. Miklukho-Maklaya 16/10, 117997 Moscow, Russia; fax: +7 (495) 335-0812; E-mail: office@ibch.ru; kgobova@gmail.com2Lobachevsky State University of Nizhni Novgorod, pr. Gagarina 23, 603950 Nizhni Novgorod, Russia
3Lomonosov Moscow State University, Faculty of Biology, 119234 Moscow, Russia
4Prokhorov General Physics Institute, Russian Academy of Sciences, ul. Vavilova 38, 119991 Moscow, Russia; fax: +7 (499) 135-0270
* To whom correspondence should be addressed.
Received July 20, 2014; Revision received September 3, 2014
Here we propose a simple and reliable approach for detection of the tumor marker HER2/neu using the targeting fluorescent hybrid protein DARPin-mCherry. As a targeting module, we used DARPin9-29, which is a member of a novel class of non-immunoglobulin targeting proteins that can highly selectively recognize the extracellular domain of the epidermal growth factor receptor HER2/neu. The red fluorescent protein mCherry was used as the detecting module. The hybrid protein DARPin-mCherry was prepared with high yield in a bacterial expression system and purified in one step by affinity chromatography. The purified protein is not prone to aggregation. The specificity of DARPin-mCherry binding with the HER2/neu tumor marker was demonstrated using confocal microscopy, flow cytofluorimetry, and surface plasmon resonance. The dissociation constant of the DARPin-mCherry protein complex with the HER2/neu receptor determined by surface plasmon resonance was calculated to be 4.5 nM. These characteristics of the hybrid protein DARPin-mCherry suggest it as a promising agent for immunofluorescent assay and an attractive alternative to antibodies and their fragments labeled with fluorescent dyes that are now used for this purpose.
KEY WORDS: DARPin, mCherry, tumor marker HER2/neuDOI: 10.1134/S0006297914120141
The most common approach to study the interaction and internalization of cellular receptors with their ligands is to conjugate antibodies specific to these receptors with fluorescent dye or radiolabel. We proposed a simple and reliable method for visualization of surface cellular antigens of tumor cells using a red fluorescent hybrid protein DARPin-mCherry. Human breast adenocarcinoma cell line SK-BR-3 was used as a test object. This cell line is characterized by increased expression of the tumor marker HER2/neu, which is a member of the epidermal growth factor receptor family HER 1-4 and is also overexpressed by cancer cells of ovaries, lungs, prostate, etc. [1]. The mechanism of carcinogenesis is the uncontrolled formation of homo- and heterodimers of HER2/neu that promotes an increase in proliferation and migration of cells, inhibition of apoptosis, neoangiogenesis, and finally results in tumor formation and metastasis [2, 3]. Therefore, women with HER2/neu-positive breast cancer have more aggressive disease which is more frequently accompanied by relapses compared to HER2/neu-negative cases. The tumor marker HER2/neu is one of the best-studied proteins of this type [4-6]; nevertheless, the functions of HER2/neu as a protooncogene, its internalization mechanisms, as well as the intracellular signal transduction are still not known in detail. Moreover, HER2/neu is the only member of the family for which the ligands are still not identified [2]. Thus, details of HER2/neu internalization are extremely important for elucidation of its role in the signal transduction pathways in the cell. Finally, since HER2/neu is a significant target for designing new antitumor compounds (based on nanoparticles, lentiviruses, etc.), their cell trafficking should be studied in detail.
Thus, we created a chemically and optically stable hybrid protein with constant composition containing the far-red fluorescent protein mCherry and the targeting non-immunoglobulin polypeptide DARPin. This approach might become an alternative to antibodies and their fragments labeled with low molecular weight fluorescent dyes or radionuclides for studying surface tumor markers.
MATERIALS AND METHODS
Preparation of genetic construct DARP-mCherry. The encoding sequence of DARPin9-29 was amplified on a template of plasmid pCG-Hnse-DARPin-d18-9-29 (kindly presented by Prof. A. Pluckthun, Zurich University) using specific primers 5′-tattccatatggacctgggtaagaaactg and 5′-cgccgaattcttgcaggatttcagccag. The resulting fragment of DNA was treated with restriction endonucleases NdeI and EcoRI (the restrictase-recognition sites are underlined in the structure of the primers) and ligated with the vector pET22b(+) that was pretreated with the same restrictases. Upon transformation of the E. coli strain XL1-Blue with the ligase mixture and selection of transformants containing the insertion, plasmid pDARP9-29 was prepared. The encoding sequence of the mCherry gene with a flexible hinge-like linker PKPSTPPGSS was removed using restrictases EcoRI and HindIII from the plasmid pIG6-4D5scFv-mCherry prepared earlier in our laboratory and ligated with the plasmid pDARP9-29 treated with the same restrictases. In the resulting plasmid pDARP-mCherry, the gene of interest is under control of an inducible promoter of phage T7 and contains one reading frame encoding sequences of genes DARPin9-29 and mCherry, as well as a hexa-histidine label in the 3′-terminal part.
Isolation of recombinant protein DARPin-mCherry. The E. coli strain BL21(DE3) was transformed using the plasmid pDARP-mCherry. A single colony was settled in 2 ml of YTPS medium (1% tryptone, 1% yeast extract, 45 mM K2HPO4, 5 mM KH2PO4, 100 mM NaCl, 2 mM MgCl2) supplemented with ampicillin (0.1 g/liter) and grown overnight with aeration at 37°C. Then the overnight culture was diluted 50-fold with YTPS medium and cultured under the same conditions up to the middle logphase (OD600 ~ 0.5-0.7), and then temperature was decreased to 25°C and isopropyl thiogalactopyranoside (IPTG) was added to the final concentration of 1 mM. Induction was performed for 8 h. Then the cells were cooled and harvested by centrifugation at 6000g for 10 min at 4°C. The cell precipitate was resuspended in buffer (200 mM NaCl, 30 mM NaH2PO4, 2 mM Tris, pH 8.3), and then the cells were sonicated on ice bath with an ultrasonic disintegrator (ultrasonication for 45 s was performed 5-7 times). The cellular debris was removed by centrifugation at 15,000g for 30 min. The supernatant was passed through a filter with pores of 0.22-µ diameter and then placed onto a column of Ni2+-NTA Sepharose (GE Healthcare, GB) according to the manufacturer’s protocol. Proteins not bound to the sorbent were removed by washing the column with buffer T2 (2 mM Tris-HCl, 200 mM NaCl, 30 mM NaH2PO4, pH 8.3). The bound protein was eluted with buffer T2 supplemented with 150 mM imidazole. The yield of the purified protein was 40 mg per liter of culture.
The purified DARPin-mCherry protein was analyzed in 12% polyacrylamide gel under denaturing conditions.
Determination of binding constant of DARPin-mCherry with receptor HER2/neu. The dissociation constant of the DARPin-mCherry protein complex with the extracellular domain of the HER2/neu receptor was determined using surface plasmon resonance with a Biacore device (GE Healthcare). CM-5 sensor chips were covered with recombinant antigen p185HER-2-ECD with density of 4500 resonance units (RU). The dissociation constant was calculated from the resulting sensogram using the BIAevaluation 3.0 program (GE Healthcare).
Confocal microscopy. The cell lines SKBR-3 and CHO were cultured in RPMI-1640 medium (Thermo Scientific, Russia) supplemented with 10% fetal calf serum (Thermo Scientific) in culture flasks. For determination of DARPin-mCherry binding to HER2/neu receptor, the cells at the density 15,000 cells/ml were plated into confocal dishes with glass bottom (WillCo Well, The Netherlands) and grown overnight at 37°C in an atmosphere with 5% CO2. Before the experiment, the cells were supplemented with protein DARPin-mCherry to final concentration 1 µM and incubated at 4°C for 30 min. Then the cells were washed thrice in phosphate buffered saline (PBS) and visualized with a Carl Zeiss LSM-710-NLO confocal microscope (Carl Zeiss, Germany) using the following parameters: excitation at 561 nm, registration of fluorescence in the range 570-735 nm.
Flow cytofluorimetry. For quantitative evaluation of DARPin-mCherry binding to SKBR-3 (HER2/neu-overexpressing) and CHO (HER2/neu-negative) cell lines, the cells were incubated in flasks and detached with Versene solution (not using trypsin). The cells were incubated with 1 µM solution of DARPin-mCherry for 15 min at 4°C, washed twice with cold PBS, and measurements were performed using a FACS Scan Calibur device (Becton-Dickinson, USA). The data were analyzed with the WinMDI 2.8 program (Scripps Research Institute).
RESULTS AND DISCUSSION
The purpose of the present work was to create a chemically and optically stable preparation that could be easily produced in a heterologous expression system, having high affinity to the target and high signal/noise ratio upon detection. For this, we used a combination of two proteins: the non-immunoglobulin protein DARPin9-29 specifically binding to the tumor marker HER2/neu [7] and the red fluorescent protein mCherry.
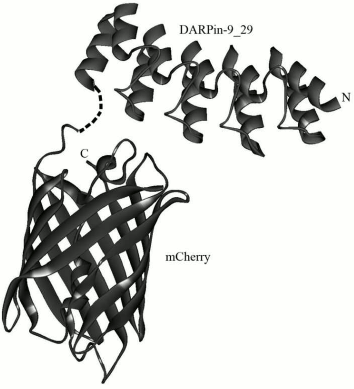
A new class of non-immunoglobulin targeting molecules called DARPins (Designed Ankyrin Repeat Proteins) was developed [8] in the laboratory of Prof. A. Pluckthun (Zurich University, Switzerland) some years ago. It was performed by selection from the library of protein ankyrin repeats using technologies of ribosomal or phage display. The structure of DARPins includes different numbers of ankyrin repeats, usually from four to six, which form a recognizing domain. One ankyrin repeat usually consists of 33 amino acid residues that form a β-turn and two antiparallel α-helices (Fig. 1). The ankyrin repeats are placed one above another and thus form a right-hand solenoid containing a longitudinal hydrophobic core and a hydrophilic surface available to the solvent [9, 10].
Fig. 1. 3D model of hybrid protein DARPin-mCherry. The figure was prepared using the DS ViewerPro 5.0 software based on DARPin9-29 (PDB accession number – 4HRL) and mCherry (accession number 2H5Q) coordinates.
High-affinity binding of these proteins to the specific target triggers different mechanisms in the cell – from inhibition of enzymes to protein–protein binding. DARPins are highly stable and have good solubility and an order of magnitude lower size compared to antibodies (14-20 kDa instead of 150 kDa), which allows a greater amount of protein to be produced in a heterologous expression system.
The pseudotyping of lentiviruses with a DARPin protein increases the specificity of their accumulation in the tumor as compared to lentiviruses pseudotyped with glycoprotein [11]. Thus, all the above-mentioned features recommend DARPins as alternatives to antibodies and mini-antibodies for application as targeting modules in hybrid constructs.
The binding and the subsequent transport of DARPins into a cell are usually quantitatively evaluated using radioactive or fluorescent labeling. However, the radioimmune assay did not give reliable results because it requires a high number of washing cycles and separation of intact cells from those with damaged membrane. Therefore, the most adequate model for study of the interaction of DARPin proteins with cells is their labeling with fluorescent dyes, e.g. with Alexa. For this, codons were introduced by directed mutagenesis providing the appearance of C- or N-terminal cysteine residues in the DARPin molecule, and then the dye was chemically linked [9].
In the present work, the protein mCherry was used as a fluorescent label, and this allowed us to avoid chemical modification of the targeting module and to prepare a chemically and optically stable sample of the desired fluorescent protein DARPin-mCherry. Being excited by long wavelength irradiation, proteins with emission in the far-red spectral region, including mCherry, are very promising as visualizing agents for animal models because they are excited by visible light. Spectral characteristics of such proteins allow researchers to separate the desired fluorescent signal from autofluorescence of the tissue due to difference in their spectral profiles [12]. mCherry is an improved form of the protein mRFP1 and has some clear advantages despite lower quantum yield. mCherry has higher extinction coefficient, is less sensitive to N- and C-terminal amino acid insertions, matures more rapidly [13, 14], and is resistant to changes in pH, which simplifies its use for conjugation with nanoparticles.
To prepare the hybrid protein DARPin-mCherry, the genetic construct pDARP-mCherry was created. Under the control of the phage T7 inducible promoter, this construct carried the gene of the recombinant protein DARP-mCherry that contained the non-immunoglobulin targeting module DARPin9-29 (possessing high affinity for the tumor marker HER2/neu), the red fluorescent protein mCherry, and also a hexahistidine sequence on the 3′-end. The C-terminal domain of the targeting module DARPin9-29 was linked with the N-terminal domain of the fluorescent module mCherry via a flexible hinge-like linker PKPSTPPGSS (Fig. 1).
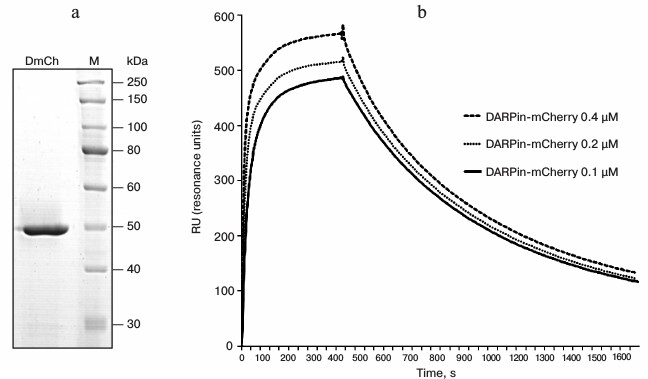
The genetic construct was expressed in the E. coli BL21(DE3) strain, and the hybrid protein DARP-mCherry was subsequently purified from the soluble fraction of cell lysate. The bright red color of protein mCherry allowed us to follow visually both its expression during cultivation of the strain-producer and the purification of the hybrid protein during metal-affinity chromatography. Analysis of the composition of the induced culture in 12% polyacrylamide gel revealed a high level of expression of the protein in soluble form, which significantly simplified the procedure of its purification. The protein was purified in one step using metal-affinity chromatography, and analysis of the resulting preparation showed the absence of other protein admixtures (Fig. 2a). The yield of the purified protein was 40 mg/liter of the liquid culture. According to electrophoretic data, the molecular weight of the protein was 46 kDa, which corresponds to the calculated value.
Fig. 2. Characteristics of hybrid protein DARPin-mCherry. a) Electrophoresis of the purified DARPin-mCherry preparation in 12% polyacrylamide gel. M, markers of molecular weight (10-250 kDa; SibEnzyme, Russia). b) Sensogram of DARPin-mCherry binding with a CM-5 sensory chip. The sensogram lines correspond to the DARPin-mCherry binding at three different concentrations with the chip covered with recombinant antigen p185HER/2-ECD (the extracellular domain of the HER2/neu receptor).
DARPin-29 was shown to interact with subdomain I of the HER2/neu receptor [15] on the tumor cell surface with an affinity of KD = 3.8 nM [7]. The dissociation constant of the hybrid DARPin-mCherry protein complex with the extracellular domain of HER2/neu was measured with a Biacore device. For this measurement, recombinant antigen p185HER2-ECD was immobilized on the surface of a CM-5 sensory chip with density of 4500 resonance units (RU). The dissociation constant was calculated from data of the sensogram (Fig. 2b) using the BIAevaluation 3.0 software program, and the value of 4.5 nM was obtained, which is insignificantly higher than the value for the free DARPin9-29 [7].
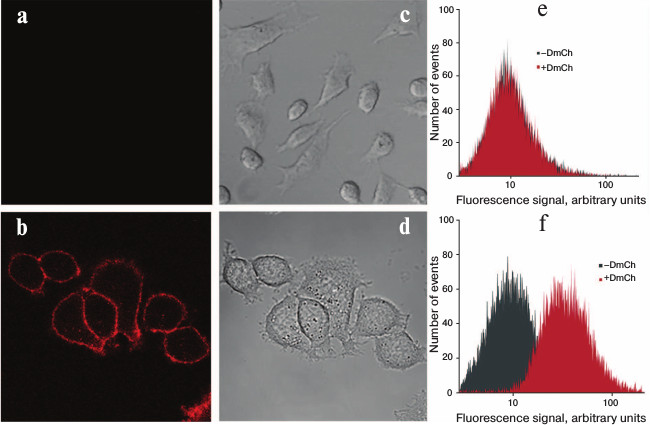
Specificity of the recombinant DARPin-mCherry protein binding with the tumor marker HER2/neu was characterized using confocal microscopy (Fig. 3). For this, the human mammary gland adenocarcinoma cell line SKBR-3 hyperexpressing the HER2/neu receptor was incubated with solution of 1 µM protein. The cells were washed thrice in phosphate–saline buffer from the unbound protein and then visualized using a Carl Zeiss LSM-710-NLO confocal microscope. The bright fluorescent signal on the surface of SKBR-3 cells indicated that the DARP-mCherry protein effectively bound to the HER2/neu receptor on their surface (Fig. 3b). To control the specificity of the DARP-mCherry binding with the HER2/neu receptor, we used the CHO cell line that does not express this tumor marker on their surface. The absence of the fluorescent signal on the surface of the CHO cells (Fig. 3a) confirmed the selective interaction of the hybrid DARPin-mCherry protein with the tumor marker HER2/neu on the surface of tumor cells.
Fig. 3. Detection of hybrid protein DARPin-mCherry binding with HER2/neu-positive cells. Confocal images are presented for CHO cells (a, c) and SKBR-3 (b, d) treated with DARPin-mCherry; images (a) and (b) obtained in the red channel of fluorescence correspond to images (c) and (d) obtained in transmitted light. Results of flow cytofluorimetry are shown for the CHO (e) and SKBR-3 cells (f) before treatment with DARPin-mCherry (plots with gray shading) and after incubation with DARPin-mCherry (plots with red shading).
Thus, the obtained hybrid protein can be used for visualization of HER2/neu-positive tumor cells. The optical properties of the protein as a label in microscopy (especially for qualitative estimation) predetermine its use in flow cytometry for detection of the HER2/neu receptor on the cell surface (quantitative estimation).
Thus, SKBR-3 cells incubated with DARPin-mCherry demonstrated significant increase in the mean intensity of fluorescence compared to the control (Fig. 3f). Such effect was not observed in the population of CHO cells.
We obtained a new fluorescent hybrid protein DARPin-mCherry, which is expressed in E. coli system with high yield and purified by metal affinity chromatography in one step. The specificity and affinity of DARPin-mCherry protein to the HER2/neu receptor have been characterized. High specificity and photostability make this protein a promising agent for immunofluorescence assay. We conclude that hybrid fluorescent proteins based on DARPins are promising agents for immunofluorescence assay and a good alternative for antibodies and their fragments [16] labeled with low molecular weight fluorescent dyes.
Since there is a repertoire of DARPins possessing high efficiency to different tumor markers [17, 18], this approach can be implemented for specific labeling of other cell surface antigens with different fluorescent proteins [14, 19].
Authors are grateful to Prof. V. B. Loshchenov (Prokhorov Institute of General Physics) for the opportunity to perform experiments using confocal microscopy.
The work was supported by the Russian Foundation for Basic Research (projects 12-04-01083a, 14-04-32156 mol_a, 13-04-40226-N KOMFI, 12-04-00757a) and by the Russian Federation Ministry of Education and Science (project 14.Z50.31.0022).
REFERENCES
1.Slamon, D. J., Godolphin, W., Jones, L. A., Holt,
J. A., Wong, S. G., Keith, D. E., Levin, W. J., Stuart, S. G., Udove,
J., Ullrich, A., and Press, M. (1989) Studies of the HER-2/neu
proto-oncogene in human breast and ovarian cancer, Science,
244, 707-712.
2.Polyanovski, O. L., Lebedenko, E. N., and Deyev, S.
M. (2012) ERBB-oncogenes as targets for monoclonal antibodies,
Biochemistry (Moscow), 77, 227-245.
3.Citri, A., Gan, J., Mosesson, Y., Vereb, G.,
Szollosi, J., and Yarden, Y. (2004) Hsp90 restrains ErbB-2/HER2
signaling by limiting heterodimer formation, EMBO Rep.,
5, 1165-1170.
4.Sorkin, A., and Goh, L. K. (2009) Endocytosis and
intracellular trafficking of ErbBs, Exp. Cell Res.,
315, 683-696.
5.Yarden, Y., and Sliwkowski, M. X. (2001) Untangling
the ErbB signaling network, Nature Rev. Mol. Cell Biol.,
2, 127-137.
6.Arteaga, C. L., and Engelman, J. A. (2014) ERBB
receptors: from oncogene discovery to basic science to mechanism-based
cancer therapeutics, Cancer Cell, 25,
282-303.
7.Steiner, D., Forrer, P., and Pluckthun, A. (2008)
Efficient selection of DARPins with sub-nanomolar affinities using SRP
phage display, J. Mol. Biol., 382, 1211-1227.
8.Binz, H. K., Stumpp, M. T., Forrer, P., Amstutz,
P., and Pluckthun, A. (2003) Designing repeat proteins: well-expressed,
soluble and stable proteins from combinatorial libraries of consensus
ankyrin repeat proteins, J. Mol. Biol., 332,
489-503.
9.Tamaskovic, R., Simon, M., Stefan, N., Schwill, M.,
and Pluckthun, A. (2012) Designed ankyrin repeat proteins (DARPins)
from research to therapy, Methods Enzymol., 503,
101-134.
10.Kobe, B., and Kajava, A. (2000) When protein
folding is simplified to protein coiling: the continuum of solenoid
protein structures, Trends Biochem. Sci., 25,
509-515.
11.Munch, R. C., Muhlebach, M. D., Schaser, T.,
Kneissl, S., Jost, C., Pluckthun, A., Cichutek, K., and Buchholz, C. J.
(2011) DARPins: an efficient targeting domain for lentiviral vectors,
Mol. Ther., 19, 686-693.
12.Borovjagin, A. V., McNally, L. R., Wang, M.,
Curiel, D. T., MacDougall, M. J., and Zinn, K. R. (2010) Noninvasive
monitoring of mRFP1- and mCherry-labeled oncolytic adenoviruses in an
orthotopic breast cancer model by spectral imaging, Mol.
Imaging, 9, 59-75.
13.Shaner, N. C., Campbell, R. E., Steinbach, P. A.,
Giepmans, B. N., Palmer, A. E., and Tsien, R. Y. (2004) Improved
monomeric red, orange and yellow fluorescent proteins derived from
Discosoma sp. red fluorescent protein, Nature
Biotechnol., 22, 1567-1572.
14.Chudakov, D. M., Matz, M. V., Lukyanov, S. A.,
and Lukyanov, K. A. (2010) Fluorescent proteins and their applications
in imaging living cells and tissues, Physiol. Rev., 90,
1103-1163.
15.Jost, C., Schilling, J., Tamaskovic, R., Schwill,
M., Honegger, A., and Pluckthun, A. (2013) Structural basis for
eliciting a cytotoxic effect in HER2-overexpressing cancer cells via
binding to the extracellular domain of HER2, Structure,
21, 1979-1991.
16.Deyev, S. M., and Lebedenko, E. N. (2009) Modern
technologies for creation of artificial antibodies for clinical
application, Acta Naturae, 1, 32-50.
17.Boersma, Y. L., Chao, G., Steiner, D., Wittrup,
K. D., and Pluckthun, A. (2011) Bispecific designed ankyrin repeat
proteins (DARPins) targeting epidermal growth factor receptor inhibit
A431 cell proliferation and receptor recycling, J. Biol. Chem.,
286, 41273-41285.
18.Souied, E. H., Devin, F., Mauget-Faysse, M.,
Kolar, P., Wolf-Schnurrbusch, U., Framme, C., Gaucher, D., Querques,
G., Stumpp, M. T., and Wolf, S. (2014) Treatment of exudative
age-related macular degeneration with a designed ankyrin repeat protein
that binds vascular endothelial growth factor: a phase I/II study,
Am. J. Ophthalmol., DOI: 10.1016/j.ajo.2014.05.037.
19.Piatkevich, K. D., and Verkhusha, V. V. (2011)
Guide to red fluorescent proteins and biosensors for flow cytometry,
Methods Cell Biol., 102, 431-461.