Characterization of Two Recombinant 3-Hexulose-6-Phosphate Synthases from the Halotolerant Obligate Methanotroph Methylomicrobium alcaliphilum 20Z
O. N. Rozova1, S. Y. But1, V. N. Khmelenina1*, A. S. Reshetnikov1, I. I. Mustakhimov2, and Y. A. Trotsenko1,2
1Laboratory of Methylotrophy, Skryabin Institute of Biochemistry and Physiology of Microorganisms, Russian Academy of Sciences, 142290 Pushchino, Moscow Region, Russia; E-mail: khmelenina@rambler.ru2Pushchino State Institute of Natural Sciences, 142290 Pushchino, Moscow Region, Russia
* To whom correspondence should be addressed.
Received September 12, 2016; Revision received October 18, 2016
Two key enzymes of the ribulose monophosphate (RuMP) cycle for formaldehyde fixation, 3-hexulose-6-phosphate synthase (HPS) and 6-phospho-3-hexulose isomerase (PHI), in the aerobic halotolerant methanotroph Methylomicrobium alcaliphilum 20Z are encoded by the genes hps and phi and the fused gene hps-phi. The recombinant enzymes HPS-His6, PHI-His6, and the two-domain protein HPS–PHI were obtained by heterologous expression in Escherichia coli and purified by affinity chromatography. PHI-His6, HPS-His6 (2 × 20 kDa), and the fused protein HPS–PHI (2 × 40 kDa) catalyzed formation of fructose 6-phosphate from formaldehyde and ribulose-5-phosphate with activities of 172 and 22 U/mg, respectively. As judged from the kcat/Km ratio, HPS-His6 had higher catalytic efficiency but lower affinity to formaldehyde compared to HPS–PHI. AMP and ADP were powerful inhibitors of both HPS and HPS–PHI activities. The two-domain HPS–PHI did not show isomerase activity, but the sequences corresponding to its HPS and PHI regions, when expressed separately, were found to produce active enzymes. Inactivation of the hps-phi fused gene did not affect the growth rate of the mutant strain. Analysis of annotated genomes revealed the separately located genes hps and phi in all the RuMP pathway methylotrophs, whereas the hps-phi fused gene occurred only in several methanotrophs and was absent in methylotrophs not growing under methane. The significance of these tandems in adaptation and biotechnological potential of methylotrophs is discussed.
KEY WORDS: methylotrophic bacteria, Methylomicrobium alcaliphilum 20Z, 3-hexulose-6-phosphate synthase, 6-phospho-3-hexulose isomerase, ribulose monophosphate cycleDOI: 10.1134/S0006297917020092
Abbreviations: FA, formaldehyde; F6P, fructose 6-phosphate; GPD, glucose-6-phosphate dehydrogenase; HPS, 3-hexulose-6-phosphate synthase; LB, Luria–Bertani medium; ORF, open reading frame; PGI, phosphoglucoisomerase; PHI, 6-phospho-3-hexulose isomerase; RuMP, ribulose monophosphate; Ru5P, ribulose 5-phosphate.
Aerobic bacteria utilizing reduced C1-compounds such as
methane and its oxidized and substituted derivatives as growth
substrates (methylotrophs) are ubiquitous in various ecosystems where
they regulate C1-compound contents. Studies of methylotrophy
as a microbial nutrition mode are essential for understanding of
evolutionary scenario of the Earth’s life, as well as for use in
biotechnologies as catalysts for production of various useful
metabolites from renewable (and commonly toxic)
C1-substrates. Methylotrophic bacteria assimilate
C1-compounds via three specific biochemical pathways, where
the key enzymes perform first formation of a C–C bond. In the
serine pathway, the primary product serine is formed via condensation
of formaldehyde and glycine catalyzed by serine transhydroxymethylase
(EC 2.1.2.1), and further phosphosugar biosynthesis proceeds by
gluconeogenesis. The key enzymes of the ribulose monophosphate (RuMP)
cycle, 3-hexulose-6-phosphate synthase (HPS, EC 4.1.2.43) and
6-phospho-3-hexulose isomerase (PHI, EC 5.3.1.27), catalyze
condensation of formaldehyde with ribulose 5-phosphate (Ru5P) and
isomerization of D-arabino-3-hexulose 6-phosphate into
D-fructose 6-phosphate [1-4].
Some aerobic methylotrophs oxidize C1-substrates into
CO2, which further is fixed via the Calvin cycle.
The RuMP cycle is the most efficient C1-assimilation pathway [5]. Despite its key role in carbon utilization, HPS has been characterized from only a few methylotrophic bacteria. The enzyme from the thermotolerant methanotroph Methylococcus capsulatus Bath was found to be a unique hexameric membrane protein [1]. Later, the genome of this bacterium was found to contain three copies of the hps genes: in the structure of the duplicated hps/phi operon and another gene hps-phi encoding HPS–PHI fused protein. As follows from analysis of the molecular mass of the enzyme, only the HPS–PHI fused protein has been characterized. The functionality, properties, and role of HPS remained beyond researchers’ vision.
Orthologs of HPS and PHI have been found in a variety of bacteria not growing on C1-substrates. In Bacillus subtilis and Burkholderia cepacia TM1, these enzymes are involved in formaldehyde detoxification, whereas in some archaea they perform pentose phosphate formation from fructose 6-phosphate, thus compensating for the absence of the pentose phosphate pathway [6-8]. In some archaea, HPS and PHI are expressed as a two-domain protein [9, 10]. It was suggested that bifunctional HPS–PHI enhances efficiency of sequential reactions and stability against thermal denaturation in comparison to the individual enzymes.
The halotolerant methanotroph Methylomicrobium alcaliphilum 20Z is a promising organism for biotechnological application for synthesis of various useful metabolites from methane or methanol because this strain demonstrates high growth rate and ability to grow in a wide range of salinity, pH, and methanol concentrations; also, it can ferment formaldehyde under oxygen-limited conditions [11, 12]. Recently, in the genome of this methanotroph putative hps and phi genes and the additional hps-phi fused gene were found. The goal of this work was to elucidate functionality and properties of the two HPSs, of the two forms of the hexulose phosphate synthase from M. alcaliphilum 20Z obtained by heterologous expression in Escherichia coli.
MATERIALS AND METHODS
Bacteria and growth conditions. Methylomicrobium alcaliphilum 20Z (VKM B-2133T = NCIMB 14124T) was grown at 30°C in a methane–air atmosphere (1 : 1) or in the presence of 0.3% methanol (v/v) on mineral 2P medium containing 0.1 M NaHCO3 and 0.3 M NaCl [11]. Escherichia coli Rosetta (DE3) obtained from Stratagene (USA) was routinely grown at 37°C in LB medium [13]. Ampicillin (100 µg/ml), kanamycin (100 µg/ml), and chloramphenicol (25 µg/ml) were added to the growth medium if required.
Cloning and expression of the hps and hps-phi genes. DNA was isolated from M. alcaliphilum 20Z as described previously [14]. The hps (CCE25609), phi (CCE25608), and hps-phi fused (CCE23598) genes from the genomic DNA were amplified using the appropriate primers (Table 1). The PCR products were digested by endonucleases NdeI and HindIII and ligated into the expression vector pET30a(+) treated with the same restrictases. The hps-phi gene was cloned into the pHUE vector [15] between the SacII and HindIII endonuclease sites. Analogously, DNA fragments containing ORFs coding for either the synthase (hps-2) or isomerase (phi-2) domain of the HPS–PHI fused protein were amplified using HPS–HPI (NdeI)F and Delta-HPS-R(HindIII) or Delta-HPI-F and Delta-HPI-R primers, respectively (Table 1). All resultant plasmids were transformed into E. coli (DE3) Rosetta cells, and protein expression was induced by adding 0.5 mM isopropyl-1-thio-β-D-galactopyranoside (Sigma-Aldrich, USA) in the logarithmic phase of the culture (OD600 of 0.6-0.8). After 18-h growth at 18°C, the cells were harvested by centrifugation for 30 min at 5000g and 4°C.
Table 1. Primers used in this work.
Restriction sites are underlined

Purification of the recombinant enzymes. The recombinant HPS, HPS-2, PHI, and PHI-2 were purified by affinity chromatography on a Ni2+-nitrilotriacetic acid (Ni-NTA) column as described [16]. A modified protocol was used to purify the recombinant HPS–PHI. The E. coli cells were suspended in 100 mM Tris-HCl buffer, pH 8.0, containing 0.8 M NaCl, 2 mM imidazole, and 5 mM MgCl2. The cells were disrupted by sonication for 1 min with 10-s bursts, with 30-s cooling on ice between bursts, using an MSE sonicator (MSE, UK), and centrifuged for 30 min at 11,000g at 4°C. The supernatant was loaded into a Ni-NTA column equilibrated with the same buffer. The column was washed with 100 mM Tris-HCl buffer (pH 8.0) containing 0.8 M NaCl, 5 mM MgCl2, and 4 mM imidazole. The ubiquitin-His6-tagged protein was eluted with the same buffer containing 150 mM imidazole. The recombinant His6-Ub-HPS–PHI was additionally treated with deubiquitinase (overnight at 4°C), dialyzed against 100 mM Tris-HCl buffer, pH 8.0, with 0.8 M NaCl, and loaded again into the Ni-NTA column equilibrated with this buffer. The protein not bound with the column was collected and stored at 4°C.
Determination of quaternary structure. The quaternary form of the enzyme was analyzed by nondenaturing gel electrophoresis using pore-limiting gradient polyacrylamide (4-30%) [17]. Ferritin (440 kDa), amylase (200 kDa), alcohol dehydrogenase (150 kDa), and bovine serum albumin (66 kDa) from Sigma-Aldrich Group were used as reference proteins.
Enzyme assays. Two methods were used to determine HPS activity. Method I based on the measurement of formaldehyde concentration was used to determine the pH and temperature optima. The reaction mixture (0.5 ml) contained 100 mM Tris-HCl buffer, pH 8.5 (for HPS–PHI), or 100 mM NaOH-glycine buffer, pH 9.5 (for HPS), 5 mM MgCl2, 2.5 mM Ru5P, and the enzyme. The mixture was incubated for 3 min at 30°C, and the reaction was started by adding 0.5 mM formaldehyde. After 10 min, 0.5 ml of the Nash reagent [18] was added, the mixture was incubated at 50°C for 10 min, and the optical density at 420 nm was measured.
In method II, PHI-His6 purified from M. alcaliphilum 20Z, phosphoglucoisomerase (PGI), and glucose-6-phosphate dehydrogenase (GPD) were used as the coupling enzymes. The reaction mixture (see method I) additionally contained 10 U of PHI, 10 U of PGI and GPD, and 0.5 mM NADP+. The rate of NADP+ reduction was assayed at 340 nm with a Shimadzu UV-1700 spectrophotometer. Method II was used to determine the kinetic parameters of the enzymes.
The apparent values Km and Amax were calculated using the Enzyme Kinetics Module for SigmaPlot 12. The following buffers were used to test the effect of pH on the activity: potassium phosphate (pH 6.0-8.0), Tris-HCl (pH 7.0-9.0), and NaOH-glycine (pH 9.0-10.5). The following compounds were tested as potential effectors: pyruvate, citrate, serine, phosphoenolpyruvate, glycerate, 2-ketoglutarate, malate, AMP, ADP, ATP, pyrophosphate, and orthophosphate at a final concentration of 1 mM; glucose, glucose 1-phosphate, glucose 6-phosphate, fructose, fructose 1-phosphate, fructose 6-phosphate (F6P), and fructose 1,6-bisphosphate at final concentration 5 mM.
The reverse reaction catalyzed by HPS was assayed by detecting formaldehyde formation from F6P with the Nash reagent. The reaction was performed in 100 mM Tris-HCl buffer, pH 8.5 (in the case of HPS–PHI) or 100 mM NaOH-glycine buffer, pH 9.5 (in the case of HPS). The mixture (0.5 ml) contained 10 mM F6P, 10 mM MgCl2, HPS (or HPS–PHI), and PHI-His6.
Mutant generation. Insertions in the hps and phi genes and the hps-phi fused gene were generated using the suicide vector pCM184 as described [19]. Briefly, 3′- and 5′-fragments of each gene to be mutated were PCR amplified using the appropriate primers (Table 1) and inserted into pCM184 upstream and downstream of the kanamycin (Km) resistance gene. The resulting plasmids were transformed into E. coli S17-1, and the resulting donor strain was mated with wild-type M. alcaliphilum 20Z in biparental mating. Mutants were selected on 0.5% methanol plates with kanamycin (100 µg/ml). Double-crossover mutants were identified by diagnostic PCR tests.
Phylogenetic analyses. The full-length amino acid sequences from the protein databases of NCBI were used for phylogenetic analyses. The sequences were aligned using the Alignment tool of the MEGA 6 software [20]. Minor corrections of the alignments were made manually. There were 208 informative positions in the final dataset. The phylogenetic tree was generated with MEGA 6 using the Neighbor-Joining method.
RESULTS
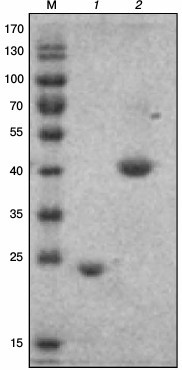
Cloning of the hps, phi, and hps-phi genes and purification of the recombinant enzymes. BLAST analysis of the M. alcaliphilum genome revealed two ORFs (CCE25609 and CCE25608) sharing noticeable sequence identities with HPS and PHI characterized from Methylomonas aminofaciens 77a (64.7 and 53.6%) and from Mycobacterium gastri MB19 (43.3 and 33.2%) [3, 8]. An additional ORF (CCE23598) coding for the putative bifunctional enzyme having both HPS and PHI domains was found. The hps and phi genes were cloned into the pET30a(+) vector designed for the expression of the C-terminal His6-tagged fusion proteins under the control of the T7 promoter. The hps-phi fused gene was cloned into the pHUE vector allowing overproduction of the recombinant enzyme with histidine-tagged ubiquitin (His6-Ub) fusion at the N-terminus [15]. The genes were expressed in E. coli Rosetta (DE3), and the recombinant enzymes were purified by affinity chromatography on Ni-NTA agarose. His6-Ub-HPS–PHI was additionally treated with a deubiquitinase to obtain the protein in its native form. The subunit molecular masses of the proteins determined by SDS-PAGE were in a good agreement with the theoretically calculated values (22.2, 19.2, and 41.9 kDa for HPS, PHI, and HPS–PHI, respectively) (Fig. 1). Nondenaturing gel electrophoresis in pore-limited gradient of acrylamide revealed single bands of about 40 kDa for HPS and PHI and one band of 70 kDa for HPS–PHI, thereby indicating that all the enzymes are homodimers (data not shown).
Fig. 1. A 12% SDS-PAGE of HPS-His6 (1) and HPS–PHI (2) from M. alcaliphilum 20Z. M, markers of molecular masses (kDa).
Kinetic parameters of recombinant HPS and HPS–PHI. Due to high instability of D-arabino-3-hexulose 6-phosphate, we were unable to characterize PHI from M. alcaliphilum 20Z. Nevertheless, the recombinant PHI demonstrated high activity and was used for measuring the HPS activity as a coupling enzyme at low concentration 0.01 mg/ml.
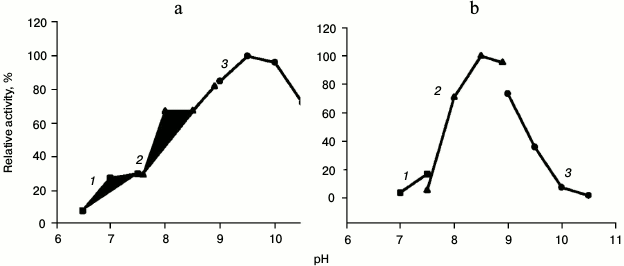
In the presence of PHI-His6, HPS-His6 catalyzed F6P formation from Ru5P and formaldehyde. The recombinant M. alcaliphilum HPS had a more alkaline pH optimum compared to the enzyme purified from M. capsulatus Bath (Fig. 2 and Table 2). This property correlated with the pH optimum for growth of M. alcaliphilum 20Z [11]. The enzyme activity was 70% higher at 50°C than at 30°C (Fig. 3). Like other studied HPSs, the M. alcaliphilum HPS was a Mg2+-dependent enzyme and strongly specific to Ru5P, since it did not use glucose, glucose 6-phosphate, glucose 1-phosphate, fructose, F6P, fructose 1,6-bisphosphate, ribose 5-phosphate, ribose 1-phosphate, or erythrose 4-phosphate as formaldehyde acceptors.
Fig. 2. Dependence of specific activity of recombinant HPS (a) and HPS-PHI (b) on pH. Curves: 1) potassium phosphate buffer; 2) Tris-HCl buffer; 3) NaOH-glycine buffer.
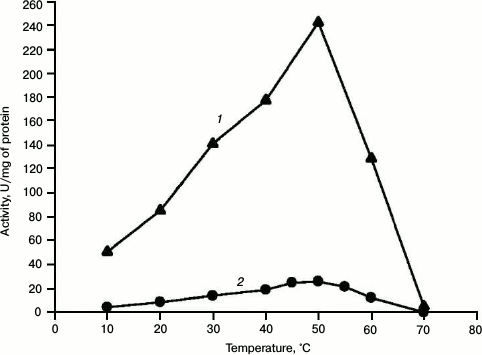
Fig. 3. Dependence of specific activity of recombinant HPS (1) and HPS–PHI fused enzyme (2) on temperature.
Table 2. Properties of hexulosephosphate
synthases from microorganisms
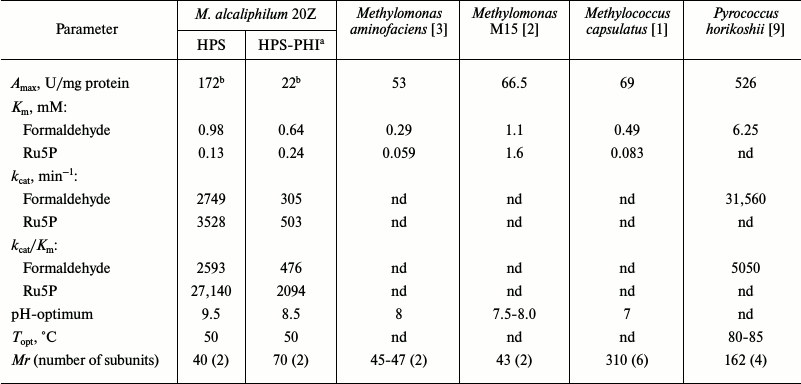
Notes: nd, not determined.
a In synthase reaction.
b Amax was determined at 30°C.
The activity of HPS was completely inhibited by 1 mM AMP. In the presence of 1 mM ADP in the reaction mixture, HPS retained 37% activity, while 1 mM ATP had only a minor inhibitory effect (Table 3). The enzyme obeyed Michaelis–Menten kinetics. Although the enzyme had a higher temperature optimum, all kinetic assays were carried out at 30°C corresponding to the optimal growth temperature for the strain. At this temperature, the maximal activity was 172 U/mg protein and the apparent Km values for formaldehyde and Ru5P were 0.98 ± 0.40 and 0.13 ± 0.01 mM, respectively. In the presence of PHI, HPS catalyzed formaldehyde formation from F6P with activity of 0.95 U/mg protein.
Table 3. Influence of some metabolites
on activity of HPS-His6 and HPS–PHI (% of activity
without effectors)
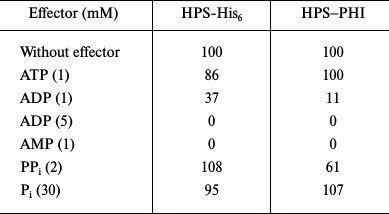
The two-domain enzyme HPS–PHI displayed HPS activity only in the presence of PHI-His6, but it showed no isomerase activity per se. The maximal activity (28 U/mg protein) was observed at 50°C and pH 8.5. The apparent Km values for formaldehyde and Ru5P measured at 30°C and pH 8.5 were 0.64 ± 0.06 and 0.24 ± 0.03 mM, respectively. Like the single-domain HPS, HPS–PHI was a Mg2+-dependent enzyme and was strongly specific to Ru5P. HPS–PHI was inhibited by ADP and completely inactivated by AMP, and it was not inhibited by ATP. In addition, PPi remarkably inhibited the enzyme activity (Table 3). The kcat/Km calculation revealed that HPS was 13-fold more efficient with Ru5P and 5-fold more efficient with formaldehyde than the HPS–PHI fused enzyme (Table 2). The reverse activity of the HPS–PHI fused enzyme (0.19 U/mg protein) was detected only in the presence of PHI, thereby confirming that the isomerase domain was unable to catalyze the reaction in either direction.
Testing of activities of HPS and PHI domains of HPS–PHI fused enzyme. We cloned the hps and phi fragments of the fused gene coding for the two-domain HPS–PHI. The His6-tagged recombinant enzymes named HPS-2 and PHI-2 were expressed in E. coli Rosetta cells and purified by one-step metal-chelating chromatography. As estimated by SDS-PAGE, the subunit molecular masses of HPS-2 and PHI-2 were ~20 kDa. Native electrophoresis showed a single band of about 40 kDa for HPS-2 and a band of 80 kDa for PHI-2 (data not shown), suggesting that the enzymes were dimer and tetramer, respectively. Notably, HPS-2 had four-fold higher synthase activity (120 U/mg protein) and two-fold higher affinity to formaldehyde (apparent Km = 0.31 ± 0.03 mM) than the HPS–PHI fused protein. Unlike the fused protein, HPS-2 obeyed Hill kinetics (n = 2.5 ± 0.6) rather than Michaelis–Menten kinetics. In contrast to HPS–PHI, the recombinant PHI-2 demonstrated isomerase activity.
Distribution and phylogenetic analysis of HPS and PHI in methylotrophs. BLAST analysis revealed the separate hps and phi genes in all gammaproteobacterial methanotrophs whose genomes are available in the database; about half have an additional hps-phi fused gene, but none of the methanotrophs has a single hps-phi fused gene (Table 4). The hps-phi fused gene is present in the moderate thermophilic and thermotolerant methanotrophs of the genera Methylocaldum and Methylococcus and halotolerant and salt-independent species of genera Methylomicrobium, Methylosarcina, and Methylovulum. The fused gene is absent in psychrophilic and salt-dependent and -independent Methylobacter and Methylomicrobium representatives, halophilic Methylohalobius, thermophilic Methylothermus, and all strains of the genus Methylomonas. Notably, the hps-phi fused gene is absent in all the RuMP pathway methylotrophic bacteria unable to use methane as growth substrate. HPS–PHI from M. alcaliphilum 20Z shared 68-99% identity of amino acid sequences with the fused enzymes from other methanotrophs, but only 38-40% identity with the two-domain enzyme from archaea of the Pyrococcus and Thermococcus genera. Interestingly, M. alcaliphilum HPS shared 73% identity with the synthase domain of its own fused enzyme. It should be noted that two or three copies of the hps and phi genes could be recognized in some representatives of methylotrophs (Table 4).
Table 4. Distribution of hps,
phi, and hps-phi fused genes in methanotrophs and
methylotrophs

Note: The database of www.genoscope.cns.fr, http://www.ncbi.nlm.nih.gov/,
and https://img.jgi.doe.gov were
used for analysis. The number of + represents the number of the
corresponding genes in the bacterial genome.
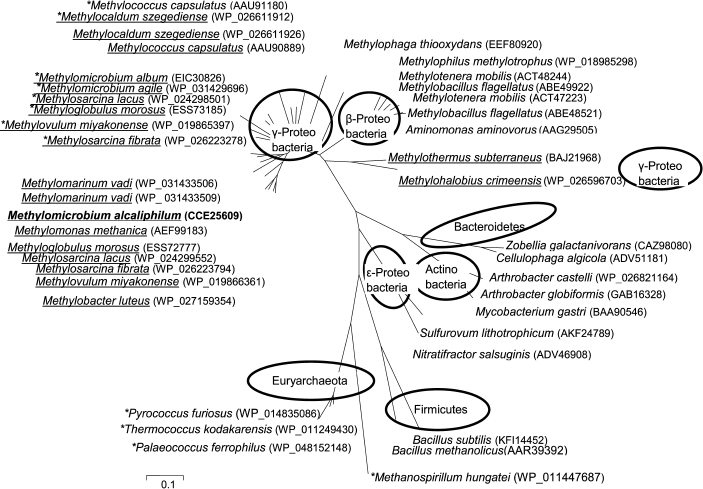
The phylogenetic analysis revealed several separate branches of HPS (Fig. 4): (1) methanotrophic Gammaproteobacteria; (2) methylobacteria of Gamma- and Betaproteobacteria classes; (3) Flavobacteroides and Actinobacteria; (4) Epsilonproteobacteria; (5) Bacillus including methylotrophs; (7) archaea. Methanotrophic HPSs are clustered together except for HPS from the halophilic Methylohalobius crimeensis and thermophilic Methylothermus subterraneus forming a separate branch on the phylogenetic tree.
Fig. 4. Phylogenetic tree of amino acid sequences of HPS and HPS–PHI fused proteins. HPS–PHI fused enzymes are marked by asterisks. Methanotrophic bacteria are underlined. Accession numbers of sequences in GenBank are indicated in parentheses.
Effect of hps-phi fused gene mutation. To elucidate the further function of the HPS–HPI fused enzyme, we constructed a hps-phi insertion mutant via allelic exchange. The growth rate of the hps-phi knockout strain was similar to that of the wild type (data not shown), most likely indicating that the HPS–PHI fused enzyme is not essential for primary carbon assimilation. In contrast, we were unable to obtain a mutant deleted in the separately located hps or phi genes.
DISCUSSION
In this work, we have first characterized two forms of HPS, which is the key enzyme of the RuMP pathway of formaldehyde fixation in aerobic methanotrophs. In M. alcaliphilum 20Z, the properties of the HPS-His6 and HPS–PHI fused enzymes suggested that the enzyme encoded by hps is responsible for assimilation of a significant part of methane carbon, whereas the two-domain protein HPS–PHI catalyzed condensation of formaldehyde and Ru5P, but with lower activity and efficiency. Moreover, the strain with an insertion in the hps-phi fused gene did not differ in growth rate from the parental strain. In addition, we were unable to obtain a viable strain with the hps or phi gene deleted.
Although HPS–PHI was inactive as isomerase, both domains became active in vitro when expressed separately, demonstrating corresponding synthase and isomerase activity. Since the active isomerase domain PHI-2 was a tetramer, while HPS–PHI has dimeric structure, we speculate that the absence of isomerase activity of the HPS–PHI fused enzyme is related to its incorrect structure. As shown, all PHI characterized to date are tetramers, and their active site is composed by all monomers of the tetramer [9, 21, 22]. Moreover, the bifunctional enzyme catalyzing the sequential HPS and PHI reactions in the archaeon Pyrococcus horikoshii OT3 had tetrameric structure [9]. The importance of quaternary structure for HPS and PHI activities was evidenced by construction of an artificial bifunctional enzyme via expression of fusions from the hps and phi genes from the methylotrophic bacterium Mycobacterium gastry without adding a linker sequence [23]. Only the product of the hps-phi fusion expressed in E. coli was active, whereas the product of phi-hps lacked activity. Since the fused enzyme from M. alcaliphilum 20Z had higher affinity to formaldehyde than the separate enzyme (Table 2), we suppose that HPS–PHI is functioning in conditions of low substrate concentrations. Much higher affinity to formaldehyde was demonstrated by the hexameric two-domain protein from M. capsulatus Bath, which also lacked isomerase activity [1].
Finally, our results can be considered in terms of molecular evolution of the RuMP pathway. Comprising separate phylogenetic groups, the sequences of the HPS and HPS domains of the fused protein of the aerobic methanotrophs are very phylogenetically remote from those of anaerobic archaea (Fig. 4). Along with high homology between HPS and the synthase domain of the fused enzyme in the same methanotroph, this could imply rather recent dividing of HPS–PHI into separate enzymes. Absence of the fused enzyme in non-methanotrophic methylotrophs sequenced to date is very intriguing.
Characterization of the key enzymes of the RuMP pathway could be applicable for genetic engineering improvement of methylotrophic producers of useful metabolites (phosphosugars, amino acids), rational fermenting with C1-compounds as auxiliary substrates, creation of transgenic plants with enhanced capability of taking up formaldehyde from air, and others [24-28]. It should be noted that Methylomicrobium buryatense 5G, the closest relative to the strain 20Z but exceeding in growth rate, has three bis-cistronic hps/phi operons in its chromosome (Table 4). While the procedure of metabolic modification is simplified when the bifunctional protein is introduced, the properties of the enzymes added demand special attention.
Acknowledgements
The authors thank all members of the Organization for Methanotroph Genome Analysis for collaboration (OMeGA) and the U.S. Department of Energy Joint Genome Institute and Genoscope for access to methanotrophic genomes for comparative analyses.
This work was supported by the Russian Science Foundation (project No. 14-14-01045).
REFERENCES
1.Ferenci, T., Strom, T., and Quayle, J. R. (1974)
Purification and properties of 3-hexulose phosphate synthase and
phospho-3-hexuloisomerase from Methylococcus capsulatus,
Biochem. J., 144, 477-486.
2.Sahm, H., Schutte, H., and Kula, M. R. (1976)
Purification and properties of 3-hexulosephosphate synthase from
Methylomonas M15, Eur. J. Biochem., 66,
591-596.
3.Kato, N., Ohashi, H., Tani, Y., and Ogata, K.
(1978) 3-Hexulosephosphate synthase from Methylomonas
aminofaciens 77a: purification, properties and kinetics,
Biochim. Biophys. Acta, 523, 238-244.
4.Arfman, N., Bystrykh, L., Govorukhina, N. I., and
Dijkhuizen, L. (1990) 3-Hexulose-6-phosphate synthase from
thermotolerant methylotroph Bacillus C1, Methods
Enzymol., 188, 391-397.
5.Quayle, J. R., and Ferenci, T. (1978) Evolutionary
aspects of autotrophy, Microbiol. Rev., 42, 251-273.
6.Yasueda, H., Kawahara, Y., and Sugimoto, S. (1999)
Bacillus subtilis yckG and yckF encode two key
enzymes of the ribulose monophosphate pathway used by methylotrophs,
and yckH is required for their expression, J. Bacteriol.,
181, 7154-7160.
7.Yurimoto, H., Hirai, R., Yasueda, H., Mitsui, R.,
Sakai, Y., and Kato, N. (2002) The ribulose monophosphate pathway
operon encoding formaldehyde fixation in a thermotolerant methylotroph,
Bacillus brevis S1, FEMS Microbiol. Lett., 214,
189-193.
8.Mitsui, R., Kusano, Y., Yurimoto, H., Sakai, Y.,
Kato, N., and Tanaka, M. (2003) Formaldehyde fixation contributes to
detoxification for growth of a nonmethylotroph, Burkholderia
cepacia TM1, on vanillic acid, Appl. Environ. Microbiol.,
69, 6128-6132.
9.Orita, I., Yurimoto, H., Hirai, R., Kawarabayasi,
Y., Sakai, Y., and Kato, N. (2005) The archaeon Pyrococcus
horikoshii possesses a bifunctional enzyme for formaldehyde
fixation via the ribulose monophosphate pathway, J. Bacteriol.,
187, 3636-3642.
10.Orita, I., Sato, T., Yurimoto, H., Kato, N.,
Atomi, H., Imanaka, T., and Sakai, Y. (2006) The ribulose monophosphate
pathway substitutes for the missing pentose phosphate pathway in the
archaeon Thermococcus kodakaraensis, J. Bacteriol.,
188, 4698-4704.
11.Khmelenina, V. N., Kalyuzhnaya, M. G.,
Sakharovsky, V. G., Suzina, N. E., Trotsenko, Y. A., and Gottschalk, G.
(1999) Osmoadaptation in halophilic and alkaliphilic methanotrophs,
Arch. Microbiol., 172, 321-329.
12.Kalyuzhnaya, M. G., Yang, S., Rozova, O. N.,
Smalley, N. E., Clubb, J., Lamb, A., Nagana Gowda, G. A., Raftery, D.,
Fu, Y., Bringel, F., Vuilleumier, S., Beck, D. A. C., Trotsenko, Y. A.,
Khmelenina, V. N., and Lidstrom, M. E. (2013) Highly efficient methane
biocatalysis revealed in methanotrophic bacterium, Nat. Commun.,
4, 2785.
13.Sambrook, J., and Russell, D. W. (2001)
Molecular Cloning: a Laboratory Manual, 3rd Edn., Cold Spring
Harbor Laboratory, N.-Y.
14.Kalyuzhnaya, M., Khmelenina, V. N., Kotelnikova,
S., Holmquist, L., Pedersen, K., and Trotsenko, Y. A. (1999)
Methylomonas scandinavica sp. nov., a new methanotrophic
psychrotrophic bacterium isolated from deep igneous rock ground water
of Sweden, Syst. Appl. Microbiol., 22,
565-572.
15.Catanzariti, A. M., Soboleva, T. A., Jans, D. A.,
Board, P. G., and Baker, R. T. (2004) An efficient system for
high-level expression and easy purification of authentic recombinant
proteins, Protein Sci., 13, 1331-1339.
16.Reshetnikov, A. S., Mustakhimov, I. I., Rozova,
O. N., Beschastny, A. P., Khmelenina, V. N., Murrell, J. C., and
Trotsenko, Y. A. (2008) Characterization of the pyrophosphate-dependent
6-phosphofructokinase from Methylococcus capsulatus Bath,
FEMS Microbiol. Lett., 288, 202-210.
17.Slater, G. G. (1969) Stable pattern formation and
determination of molecular size by pore-limit electrophoresis, Anal.
Chem., 41, 1039-1041.
18.Nash, T. (1953) The colorimetric estimation of
formaldehyde by means of the Hantsch reaction, Biochem. J.,
55, 416-421.
19.Mustakhimov, I. I., Reshetnikov, A. S., Glukhov,
A. S., Khmelenina, V. N., Kalyuzhnaya, M. G., and Trotsenko, Y. A.
(2010) Identification and characterization of EctR1, a new
transcriptional regulator of the ectoine biosynthesis genes in the
halotolerant methanotroph Methylomicrobium alcaliphilum 20Z,
J. Bacteriol., 192, 410-417.
20.Tamura, K., Dudley, J., Nei, M., and Kumar, S.
(2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software
version 4.0, Mol. Biol. Evol., 24, 1596-1599.
21.Martinez-Cruz, L. A., Dreyer, M. K., Boisvert, D.
C., Yokota, H., Martinez-Chanter, M. L., Kim, R., and Kim, S. H. (2002)
Crystal structure of MJ1247 protein from M. jannaschii at 2.0 A
resolution infers a molecular function of 3-hexulose-6-phosphate
isomerase, Structure, 10, 195-204.
22.Sanishvili, R., Wu, R., Kim, D. E., Watson, J.
D., Collart, F., and Joachimiak, A. (2004) Crystal structure of
Bacillus subtilis YckF: structural and functional evolution,
J. Struct. Biol., 148, 98-109.
23.Orita, I., Sakamoto, N., Kato, N., Yurimoto, H.,
and Sakai, Y. (2007) Bifunctional enzyme fusion of
3-hexulose-6-phosphate synthase and 6-phospho-3-hexuloisomerase,
Appl. Microbiol. Biotechnol., 76, 439-445.
24.Chen, L. M., Li, K. Z., Orita, I., Yurimoto, H.,
Sakai, Y., Kato, N., and Izui, K. (2004) Enhancement of plant tolerance
to formaldehyde by over-expression of formaldehyde-assimilating enzymes
from a methylotrophic bacterium, Plant Cell. Physiol.,
45, S233.
25.Jakobsen, O. M., Benichou, A., Flickinger, M. C.,
Ellingsen, V. S., and Brautaset, T. E. (2006) Upregulated transcription
of plasmid and chromosomal ribulose monophosphate pathway genes is
critical for methanol assimilation rate and methanol tolerance in the
methylotrophic bacterium Bacillus methanolicus, J.
Bacteriol., 188, 3063-3072.
26.Sawada, A., Oyabu, T., Chen, L. M., Li, K. Z.,
Hirai, N., Yurimoto, H., Orita, I., Sakai, Y., Kato, N., and Izui, K.
(2007) Purification capability of tobacco transformed with enzymes from
a methylotrophic bacterium for formaldehyde, Int. J.
Phytoremediat., 9, 487-496.
27.Yurimoto, H., Kato, N., and Sakai, Y. (2009)
Genomic organization and biochemistry of the ribulose monophosphate
pathway and its application in biotechnology, Appl. Microbiol.
Biotechnol., 84, 407-416.
28.Koopman, F. W., De Winde, J. H., and
Ruijssenaars, H. J. (2009) C(1) compounds as auxiliary substrate for
engineered Pseudomonas putida S12, Appl. Microbiol.
Biotechnol., 83, 705-713.