Polysaccharide Galactan Inhibits Pseudomonas aeruginosa Biofilm Formation but Protects Pre-formed Biofilms from Antibiotics
A. V. Grishin1,2,a* and A. S. Karyagina1,2,3,b*
1N. F. Gamaleya National Research Center of Epidemiology and Microbiology, Ministry of Health of the Russian Federation, 123098 Moscow, Russia2All-Russia Research Institute of Agricultural Biotechnology, 127550 Moscow, Russia
3A. N. Belozersky Institute of Physico-Chemical Biology, Lomonosov Moscow State University, 119992 Moscow, Russia
* To whom correspondence should be addressed.
Received October 9, 2018; Revised January 21, 2019; Accepted January 28, 2019
Microorganisms residing within a biofilm become more tolerant to antibiotics and other types of adverse impact, and biofilm formation by pathogenic bacteria is an important problem of current medicine. Polysaccharides that prevent biofilm formation are among the promising candidates to help tackle this problem. Earlier we demonstrated the ability of a potato polysaccharide galactan to inhibit biofilm formation by a Pseudomonas aeruginosa clinical isolate. Here we investigate the effect of potato galactan on P. aeruginosa biofilms in more detail. Microscopic analysis indicated that the galactan did not interfere with the adhesion of bacterial cells to the substrate but prevented the build-up of bacterial biomass. Moreover, the galactan not only inhibited biofilm formation, but partially destroyed pre-formed biofilms. Presumably, this activity of the galactan was due to the excessive aggregation of bacterial cells, which prohibited the formation and maintenance of proper biofilm architecture, or due to some other mechanisms of biofilm structure remodeling. This led to an unexpected effect, i.e., P. aeruginosa biofilms treated with an antibiotic and the galactan retained more viable bacterial cells compared to biofilms treated with the antibiotic alone. Galactan is the first polysaccharide demonstrated to exert such effect on bacterial biofilms.
KEY WORDS: biofilm, galactan, Pseudomonas aeruginosa, polysaccharide, antibiotics, toleranceDOI: 10.1134/S0006297919050055
Abbreviations: CBD, Calgary biofilm device; CFU, colony-forming unit; MBC, minimum bactericidal concentration; MBEC, minimum biofilm eradication concentration; MIC, minimum inhibitory concentration; PAO1, Pseudomonas aeruginosa PAO1 laboratory strain.
Biofilms are complex microbial communities enclosed in a self-produced
matrix and typically associated with a surface [1].
The process of biofilm formation is usually characterized by several
stages, namely: initial attachment of microbial cells to the surface,
aggregation of the attached cells and formation of microcolonies,
growth, maturation of the biofilm, and dispersal that allows further
dissemination of bacteria [1]. An important feature
of biofilms is their increased levels of resistance to antibiotics,
disinfectants, and the host immune system [2-5]. When biofilms of pathogenic bacteria are formed in
a human organism, they can cause such chronic conditions as infective
endocarditis, chronic wound infections, chronic otitis media, caries
and periodontitis, osteomyelitis, urinary tract infections, and chronic
lung infection in cystic fibrosis patients. Furthermore, implanted
devices and catheters are particularly prone to becoming a biofilm
substratum, i.e., the surface where biofilms form [2, 3, 6]. The
search for compounds capable of promoting biofilm dispersal or
inhibiting their formation is a growing area of research.
Polysaccharides are one of the promising classes of such compounds. In 2006, it was first reported that certain Escherichia coli capsular polysaccharides inhibited biofilm formation of several pathogenic bacterial species [7]. Since then, antibiofilm properties were demonstrated for several other polysaccharides, mostly ones of bacterial origin [8-21]. Biofilm-inhibiting properties were also shown for the plant polysaccharides galactan and galactomannan [22], and yeast- [23] and diatom-produced polysaccharides [24]. The potential advantages of polysaccharides as antibiofilm agents are low cost, biodegradability, and biocompatibility [25, 26], and further exploration of polysaccharides that disperse biofilms or inhibit their formation is an important task.
Different antibiofilm polysaccharides exert different effects on biofilms. Some polysaccharides, like PAM galactan isolated from Kingella kingae culture, Pseudoalteromonas haloplanktis-produced polysaccharide, or Pseudomonas stutzeri exopolysaccharide EPS273 can fully or partially disperse pre-formed biofilms [12, 14, 20]. Other polysaccharides only inhibit the formation of biofilms, having no effect on mature biofilms [7, 19, 21]. Apparently, the mechanism of action of antibiofilm polysaccharides is most often connected to the inhibition of initial bacterial adhesion to the substratum [10, 14, 19-21]. In some cases, polysaccharides also prevented cell–cell aggregation [7, 8, 27, 28]. Authors usually explain this effect by the alteration of physical and chemical properties of bacterial cells and/or substratum due to the interaction with a polysaccharide.
If polysaccharides are to be applied as a treatment for biofilm-associated bacterial infections, they will most probably be used in combination with traditional antibiotics. Nevertheless, only one work reported the combined use of antibiotics and a biofilm-dispersing polysaccharide. Exopolysaccharide A101 extracted from Vibrio spp. QY101 culture increased the efficacy of aminoglycoside antibiotics against P. aeruginosa biofilms by a factor of 32 [8].
Earlier, we provided evidence that potato polysaccharide galactan inhibited biofilm formation by P. aeruginosa clinical isolate 216 during 5-h cultivation [22]. The aim of this study was to further investigate the antibiofilm properties of potato galactan, including its effect on the efficacy of antibiotics commonly used to treat P. aeruginosa infections.
MATERIALS AND METHODS
Strains and media. Pseudomonas aeruginosa laboratory strain PAO1 and P. aeruginosa clinical isolate 216 were used in this work. Isolate 216 was previously isolated from a cystic fibrosis patient by the laboratory of Prof. I. A. Shaginyan (N. F. Gamaleya National Research Center for Epidemiology and Microbiology, Moscow, Russia). For long term storage, the strains were kept frozen in 40% glycerol at –80°C. During work, the strains were routinely maintained on Difco nutrient broth (Becton Dickinson, USA) with 1.5% agar. Difco nutrient broth with 1.5% agar and 1.0% NaCl was used for CFU (colony-forming unit) counting and minimum bactericidal concentration (MBC) determination. M63 medium (3 g/liter KH2PO4, 7 g/liter K2HPO4 and 2 g/liter (NH4)2SO4) supplemented with 1 mM MgSO4, 0.5% casamino acids, and 0.2% glucose [29] was used for biofilm cultivation and other experiments unless noted otherwise.
Biofilm cultivation, inhibition, and dispersion. Potato galactan (P-GALPOT) and barley glucan (P-BGBL) polysaccharides were obtained from Megazyme Inc. (Ireland). The polysaccharides were dissolved in sterile water at a concentration of 10 mg/ml. To this end, sterile water was added to dry galactan powder, and the slurry was placed on a hotplate and constantly stirred at 60°C until dissolution of the galactan. Similarly, sterile water was added to dry glucan powder, the slurry was placed on a hotplate and brought to a boil while stirring constantly, then the heating was turned off and the stirring continued until the complete dissolution of the glucan. The solutions were cooled to room temperature, and the polysaccharide concentration was brought to 10 mg/ml with sterile water. To remove any undissolved impurities, the solutions were centrifuged at 12,000g for 10 min. Then the supernatant was incubated at 80°C for 2 h. Sterility of the polysaccharide solutions was confirmed by plating on Difco nutrient broth with 1.5% agar (Becton Dickinson) and Brain Heart Infusion agar (Sifin Diagnostics, Germany) and by inoculation into Fluid Thioglycolate Medium (HiMedia Laboratories, India).
In the present work, P. aeruginosa biofilms were cultivated either on the walls of the wells of 96-well plates or on polypropylene coupons. To cultivate the biofilms on the walls of the wells of 96-well plates, P. aeruginosa PAO1 or isolate 216 were grown on nutrient agar plates for 20-24 h, then the bacteria were suspended in M63 medium supplemented with magnesium, casamino acids, and glucose to an optical density of 0.5 McFarland and diluted to the final concentration of ~(3-5)·107 CFU/ml. The resulting bacterial suspension was transferred into the 96-well plates (90 μl per well) (Costar 3599 tissue culture treated flat-bottom plates; Corning, USA), followed by 10 μl of polysaccharide solution or water (control). The plate was sealed with parafilm and incubated at 36°C for 24 h. After incubation, the plates were carefully washed using the following procedure. First, 100 μl of sterile M63 base medium was added to the plate wells, and the content of the wells were mixed by pipetting and aspirated using a multichannel pipette. Then 200 μl of M63 base medium was added to the wells and removed by inverting the plate over an autoclavable container; this procedure was performed three times. The plate was tapped on a stack of paper towels to remove the remainder of the liquid and was left in upside-down orientation at room temperature to dry overnight. To stain the biofilms, 200 μl of 0.1% crystal violet solution was added to the wells, incubated for 20-30 min, then the plate was washed with water and dried in upside-down orientation. To solubilize the biofilm-bound stain, 100 μl of 30% acetic acid was added to the wells and incubated for 20-30 min. The contents of the well were mixed by pipetting, and 90 μl of solubilized stain was transferred to another flat-bottom 96-well plate for measurements. The optical density was measured at 550 nm in a Multiscan FC plate reader (Thermo Fischer Scientific, USA). For the biofilm dispersion experiments, P. aeruginosa biofilms were cultivated in 96-well plates for 24 h and washed. Then, 90 μl of fresh M63 medium supplemented with magnesium, casamino acids, and glucose was added to the wells with the biofilms, followed by 10 μl of polysaccharides or water (control). The wells with control 24-h biofilms were left empty. The plate was sealed with parafilm and incubated at 36°C for 24 h. After that, the biofilms were washed, stained, and quantified as described above.
To cultivate P. aeruginosa biofilms on polypropylene coupons, coupons ~10.0 × 6.5 mm in size were cut out from a polypropylene sheet (Trans G03, thickness 0.5 mm; Carolex, France), sterilized by soaking in 70% ethanol, and dried. The coupons were placed vertically into the wells of 96-well plates containing 100 μl of bacterial suspension in M63 medium supplemented with magnesium, casamino acids, and glucose, with or without the galactan. The plate was sealed with parafilm and incubated at 36°C for 24 h. After incubation, the coupons were taken out from the plate wells, carefully washed by flushing with M63 base medium, dried, stained with 0.1% crystal violet for 20-30 min, washed with water, and dried again. For the biofilm dispersion experiments, the biofilms were cultivated on polypropylene coupons for 24 h, washed with sterile M63 base medium, and placed into the wells of a 96-well plate containing 100 μl of fresh medium with or without the galactan. The plate was sealed with parafilm and incubated for 24 h. After that, the coupons were taken out, washed, and stained. Since biofilms form on both sides of the coupons, the side that was oriented downward during drying was wiped with 70% ethanol to remove the biofilm before photographing and microscopy.
To quantify viable cells in biofilms, the biofilms cultivated on polypropylene coupons were used. Coupons with biofilms were placed into 1.5-ml microcentrifuge tubes containing ~500 μl glass beads Ø0.6-0.8 mm and 400 μl of M63 base medium (PAO1) or PBS (isolate 216), and vigorously vortexed for 15 s (3500 rpm, Microspin FV-2400; Biosan, Latvia). The viable cells detached from the coupons were quantified by plating and counting the colonies. It must be noted that when this bead-beating procedure was applied to planktonic cells, it caused a drop in viable cell counts by ~40-50% for PAO1 and ~10% for isolate 216. However, this decrease is considerably smaller than the difference in viable cell counts between differently treated biofilm variants, especially between intact and antibiotic-treated biofilms.
MIC, MBC, and MBEC determination. Ceftazidime and amikacin sulfate (Sintez Ltd, Russia) were freshly dissolved in M63 medium before each experiment. Working solutions of ciprofloxacin (Elfa Laboratories, India) were prepared from 2 mg/ml stock before each experiment.
Bacteria were grown on nutrient agar plates for 20-24 h, suspended in M63 medium supplemented with magnesium, casamino acids, and glucose at optical density of 0.5 McFarland, and diluted to final concentration of ~5·105 or ~5·107 CFU/ml (see “Results”). The resulting bacterial suspension was transferred into a 96-well plate (90 μl per well), followed by 10 μl of serial 2-fold dilutions of antibiotics prepared in the same medium. The plate was sealed with parafilm and incubated at 36°C for 24 h. Minimum inhibitory concentration (MIC) was then determined as the lowest concentration of an antibiotic that resulted in the absence of visible turbidity in the well plates. After MIC determination, 3 μl of the well contents was transferred to agar plates and incubated at 36°C for 24 h. Minimum bactericidal concentration (MBC) was determined as the minimum concentration of an antibiotic that resulted in the absence of growth on a nutrient agar plate. This protocol is somewhat different from the standard MBC determination procedure and instead follows the protocol described by T.-F. Mah [30].
Minimum biofilm eradicating concentrations (MBEC) of antibiotics were determined using the Calgary Biofilm Device (CBD) (MBEC Biofilm Inoculator; Innovotech Inc., Canada). The CBD is a special type of 96-well plate lid equipped with pegs that fit into the plate wells. When a CBD is used for biofilm cultivation, the biofilms form on the pegs. To cultivate the biofilms on a CBD, the CBD was placed on a 96-well plate containing 150 µl of bacterial suspension (~(3-5)·107 CFU/ml) and incubated at 36°C for 24 h. The biofilms formed on the CBD pegs were washed twice by transferring the CBD to a new plate containing 200 µl of sterile M63 base medium and shaking at 250 rpm for 1 min. Then the CBD was transferred to another 96-well plate containing 200 μl of serial 2-fold antibiotic dilutions in M63 medium supplemented with magnesium, casamino acids, and glucose, with or without the galactan, and incubated at 36°C for 24 h. After antibiotic treatment, the biofilms of the CBD pegs were washed twice, and the CBD was transferred to a plate containing 200 μl of Difco nutrient broth and incubated for 24 h to allow the surviving bacteria to resume growth. Bacterial growth was determined by the appearance of visible turbidity in the plate wells and by plating the contents of the wells onto agar plates. MBEC was determined as the minimum concentration of an antibiotic after which the biofilm was unable to resume bacterial growth.
Determination of bacterial counts in antibiotic-treated biofilms. To quantify the viable cells in the antibiotic-treated biofilms, the biofilms cultivated on polypropylene coupons were used. The biofilms were cultivated on polypropylene coupons for 24 h, washed, and placed into 1.5-ml microcentrifuge tubes containing 700 μl of the antibiotic solution in M63 medium supplemented with magnesium, casamino acids, and glucose, with or without the galactan. The tubes with coupons were incubated at 36°C for 24 h. After incubation, the viable cells within biofilms were quantified as described above. The colonies were counted in 10-μl spots on agar plates, which translates into the theoretical detection limit of ~100 CFU/ml or ~40 CFU/coupon (1.6 log10 CFU/coupon). To determine if there are any viable bacteria in the biofilms below the detection limit, 400 μl of fresh Difco nutrient broth was added to the tubes where biofilms had been disrupted, and the tubes were incubated at 36°C for 24 h. Bacterial growth was then determined by visible turbidity and plating on agar plates.
Fluorescence microscopy. PAO1 biofilms were cultivated on polypropylene coupons and treated with 256 μg/ml amikacin with or without 1 mg/ml galactan. After treatment, the biofilms were washed with water and stained with FilmTracer LIVE/DEAD Biofilm Viability Kit (Molecular Probes Inc., USA) for 15 min as per the manufacturer’s recommendations. The stained biofilms were visualized using an EVOS FLoid cell imaging station (Thermo Fischer Scientific). For the clarity of representation, pictures were edited with Fiji software [31] as follows. First, “subtract background” function with the option “create background” was used to calculate the mean background intensities for green and red channels individually. Then, red and green channels of all images were scaled to make the mean background values equal. Next, the “subtract background” function was used to remove the background, and the channels were merged to obtain the final pictures.
Statistical analysis. Statistical analysis was performed in Microsoft Excel using Real Statistics Resource Pack Excel add-on (Release 4.7, Charles Zaiontz, www.real-statistics.com). Normality of the data was assessed by the Shapiro–Wilk test. Two-tailed t-test with unequal variances or two-tailed Mann–Whitney test were used to determine the statistical significance of the differences between the group means. The differences were considered significant if p < 0.05 after applying Dunn-Šidák correction for multiple comparisons. The CFU counts were log-transformed before calculating statistical parameters and applying statistical tests. In all figures except Fig. 3, results from a single experiment are shown. However, all experiments except disruption of 48-h biofilms (Fig. 1c) were performed at least twice. In Fig. 3, results from the two latest experiments were pooled to increase the number of replicates and statistical power of the analysis. The experimental data are provided in the supplementary material [see Supplement to this paper on the website of the journal (http://protein.bio.msu.ru/biokhimiya) and Springer site (Link.springer.com)].
RESULTS
Potato galactan inhibits biofilm formation and partially disperses pre-formed biofilms. We demonstrated previously that potato galactan inhibited early stages of biofilm formation by P. aeruginosa clinical isolate 216 [22]. To further explore the effect of this galactan, we chose laboratory strain PAO1, which is widely used in P. aeruginosa biofilm studies.
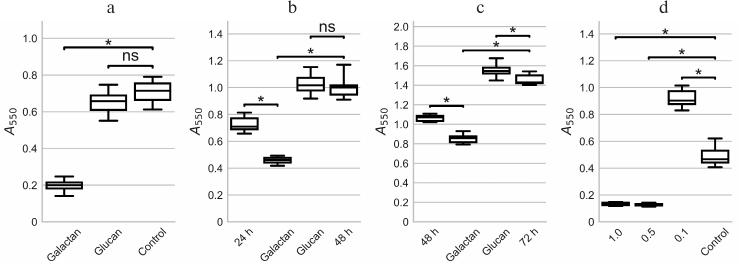
Potato galactan at a concentration of 1 mg/ml almost completely inhibited the formation of biofilms by PAO1 during 24 h of cultivation. The amount of biomass in the biofilms grown in presence of the galactan was 3.5-fold lower than in control biofilms (Fig. 1a). Barley glucan used as a control polysaccharide did not significantly influence biofilm formation (Fig. 1a). We also noted that the galactan caused intensive aggregation of bacterial cells: large slimy aggregates not attached to the plate wells were seen by the unaided eye during washing of the biofilms. These aggregates were absent in the control or glucan wells.
Fig. 1. Influence of potato galactan on formation and dispersion of P. aeruginosa PAO1 biofilms in 96-well plates. a) Formation of PAO1 biofilms in the presence of 1 mg/ml potato galactan or glucan, or without polysaccharide (Control) for 24 h. b) Dispersion of PAO1 biofilms pre-cultivated for 24 h (24 h) by 1 mg/ml potato galactan or barley glucan, or without polysaccharide (48 h). c) Dispersion of PAO1 biofilms pre-cultivated for 48 h by 1 mg/ml potato galactan or barley glucan, or without polysaccharide (72 h). d) Formation of PAO1 biofilms in presence of 1.0, 0.5, and 0.1 mg/ml galactan, or without the galactan (Control) for 24 h. The data are presented as box-plots where the box indicates 25th and 75th percentiles, the middle line indicates median value, and the whiskers show minimal and maximal values. Statistical significance of the differences between group means was determined by two-tailed t-test with unequal variances. The differences were considered significant (*) if p < 0.05 after correction for multiple comparisons; ns, non-significant.
When biofilms were first pre-cultivated for 24 h and then treated with 1 mg/ml potato galactan, partial biofilm dispersion was observed. The amount of biomass decreased by almost 40% (Fig. 1b). Control polysaccharide barley glucan did not have any effect on pre-formed biofilms (Fig. 1b). A similar pattern was seen when biofilms were treated with the galactan after 48 h of pre-cultivation (Fig. 1c).
To investigate the influence of potato galactan concentration on its antibiofilm effect, PAO1 biofilms were cultivated in the presence of 0.5 and 0.1 mg/ml of the galactan. Interestingly, although 0.5 mg/ml of the galactan acted similarly to 1 mg/ml, a 10-fold lower concentration of 0.1 mg/ml not only lacked any inhibitory effect, but rather had a significant stimulatory effect on the biofilm formation (Fig. 1d). We used the galactan at the concentration of 1 mg/ml in all further experiments.
Polymeric structure of the galactan was necessary for its antibiofilm activity, since monomeric galactose in equivalent concentration had no influence on PAO1 biofilm formation (data not shown).
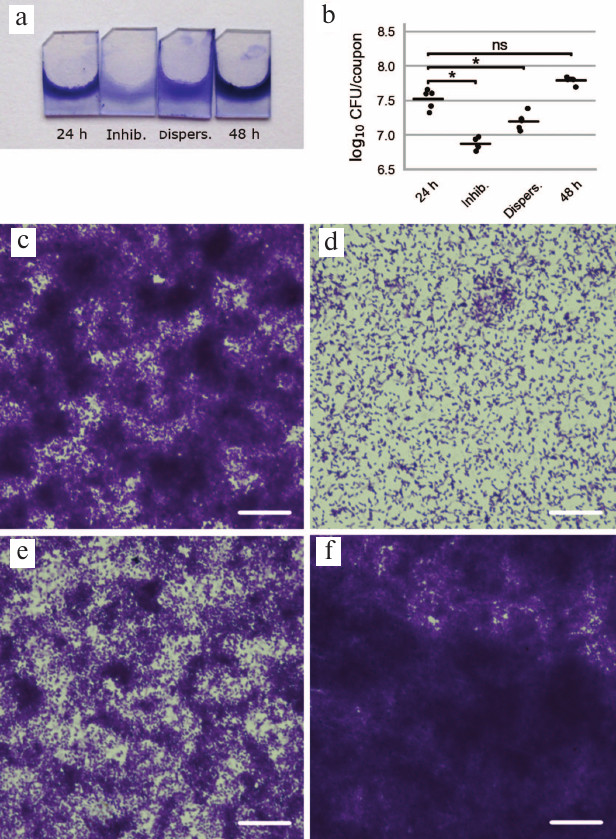
We further studied the effect of potato galactan on P. aeruginosa biofilms by microscopy. To this end, biofilms were cultivated on polypropylene coupons vertically placed into the wells of 96-well plates, dried, and stained with crystal violet. Visually, biofilm formation on polypropylene coupons followed the same pattern as the 96-well plates biofilms. Specifically, biofilm formation in the presence of the galactan was virtually abolished, and treatment of pre-formed biofilms with the galactan led to their partial disruption (Fig. 2, a and b). Microscopic analysis demonstrated that control biofilms cultivated for 24 h without galactan consisted of large and dense bacterial aggregates interspersed with more loosely packed bacteria and bacteria-free polypropylene regions (Fig. 2c). Biofilms cultivated in presence of the galactan were drastically different and showed evenly spaced bacterial cells that did not form pronounced clusters or aggregates (Fig. 2d). Galactan-treated pre-formed biofilms were similar in appearance to control biofilms but thinner; dense bacterial aggregates were smaller and occupied less area (Fig. 2e). Finally, control biofilms cultivated in the absence of the galactan for 48 h constituted a robust layer of bacterial biomass with morphological features hardly discernable by light microscopy (Fig. 2f).
Fig. 2. Influence of potato galactan on formation and dispersion of P. aeruginosa PAO1 biofilms on polypropylene coupons. a) Pictures of polypropylene coupons with biofilms stained with crystal violet (24 h – control biofilm cultivated for 24 h; Inhib. – biofilm cultivated for 24 h in presence of 1 mg/ml potato galactan; Dispers. – 24-h pre-formed biofilm treated with 1 mg/ml potato galactan for additional 24 h; 48 h – control biofilm cultivated for 48 h). b) Viable cell counts in biofilms cultivated on polypropylene coupons (designations as in (a)); data are presented as individual datapoints (n = 5), horizontal lines indicate mean values. Statistical significance of differences between group means was determined by two-tailed t-test with unequal variances. Differences were considered significant (*) if p < 0.05 after correction for multiple comparisons; ns, non-significant. c-f) Microscopic pictures of biofilms cultivated on polypropylene coupons: c) control 24-h biofilm; d) biofilm cultivated in presence of 1 mg/ml potato galactan; e) 24-h pre-formed biofilm treated with 1 mg/ml potato galactan; f) control 48-h biofilm. The scale bar is 25 μm.
Viable cell counts in biofilms cultivated on polypropylene coupons were in good agreement with the results of the biofilm biomass quantification in 96-well plates. Biofilms grown in the presence of the galactan contained approximately 4.5 times fewer viable bacterial cells than the control biofilms. Similarly, biofilms pre-cultivated for 24 h and treated with the galactan had twice fewer viable bacteria than the untreated biofilms (Fig. 2b).
Potato galactan protects biofilm-dwelling bacteria from antibiotics. The most probable scenario of how antibiofilm compounds could be applied assumes their use as a supplement to the standard antibiotic therapy. Thus, we further studied the combined effect of antibiotics and galactan on P. aeruginosa biofilms. In this work, we selected three different antibiotics of different classes routinely used in clinical practice to treat P. aeruginosa infections. These were ceftazidime (a cephalosporin), ciprofloxacin (a fluoroquinolone), and amikacin (an aminoglycoside).
The minimum inhibitory concentrations and minimum bactericidal concentrations (MICs and MBCs) of these antibiotics against PAO1 determined in this work are presented in the table. Some antibiotics are known to display so-called “inoculum effect”, when the MIC of the antibiotic increases dramatically with an increase in the initial bacterial density (i.e., the inoculum, normally 5·105 CFU/ml). The number of bacteria inside biofilms is much higher than 5·105 CFU/ml (see Fig. 2b), and if resistance to an antibiotic is observed for biofilm-dwelling bacteria, it may be caused by the inoculum effect and not by biofilm-specific mechanisms. We thus determined MICs of all three antibiotics for both standard (5·105 CFU/ml) and increased (5·107 CFU/ml) inoculum sizes (table). Ceftazidime demonstrated a pronounced inoculum effect, having MIC of 2 μg/ml with standard initial bacterial density, and >16 μg/ml in case of increased inoculum size. We thus excluded ceftazidime from further study. MICs of the two other antibiotics did not show marked dependence on the initial bacterial density. It must be noted that the MIC of amikacin was 32 μg/ml, which is a rather high value. However, this effect was likely due to the medium used (M63), since disk diffusion assay on standard Muller–Hinton agar demonstrated susceptibility of PAO1 to amikacin.
MIC, MBC, and MBEC values of antibiotics against P. aeruginosa
PAO1 and isolate 216
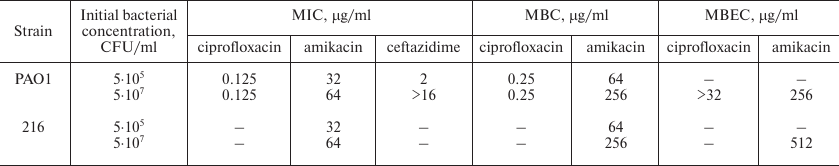
To determine the minimum biofilm eradication concentrations (MBECs), biofilms were first cultured on the pegs of the Calgary Biofilm Device (CBD) and then incubated in the antibiotic solutions. After that, the biofilms were transferred into fresh nutrient broth to allow the surviving bacteria to resume growth. The minimum concentration of amikacin that prevented the renewal of bacterial growth by PAO1 biofilms was 256 μg/ml. Ciprofloxacin MBEC was higher than the maximal tested concentration (32 μg/ml, a concentration 256 times higher than the MIC) and thus was not determined (table).
Despite the ability of the potato galactan to partially destroy PAO1 biofilms, the addition of 1 mg/ml of the galactan to the antibiotic solutions did not alter MBEC of any of the antibiotics.
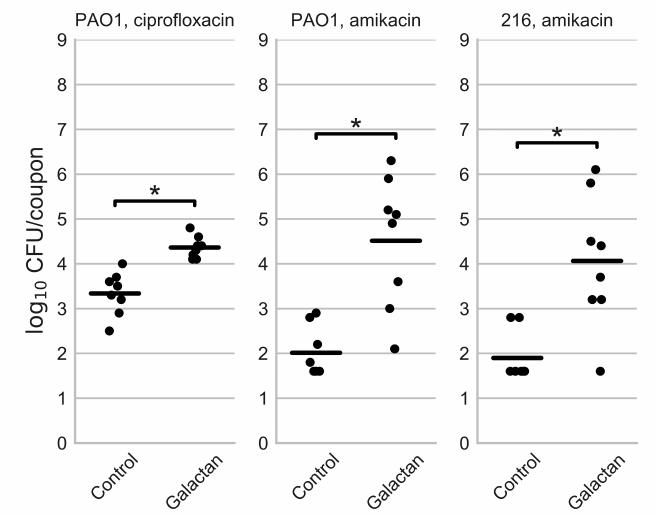
Although the galactan did not influence the MBECs of the antibiotics, we hypothesized that it might reduce the number of surviving bacteria within biofilms compared to treatment with the antibiotics alone. To test this hypothesis, we cultivated the biofilms on polypropylene coupons and incubated them in ciprofloxacin (1 μg/ml) or amikacin (256 µg/ml) solutions, with or without 1 mg/ml potato galactan. The viable cell counts in the biofilms treated this way are presented in Fig. 3.
Fig. 3. Viable cell counts in biofilms of P. aeruginosa PAO1 and isolate 216 after treatment with an antibiotic (Control) or an antibiotic in combination with potato galactan. The data are presented as individual datapoints (n = 8), horizontal lines indicate mean values. The values below detection limit are set to 1.6 log10 CFU/coupon because the biofilms disrupted by glass beads resumed bacterial growth after the addition of fresh nutrient broth, despite the absence of colonies on agar plates. Statistical significance of the differences between group means was determined by two-tailed Mann–Whitney test. The differences were considered significant (*) if p < 0.05 after correction for multiple comparisons.
Contrary to our expectations, the addition of potato galactan did not reduce the number of viable bacteria in antibiotic-treated biofilm but instead increased it. The average number of viable cells in ciprofloxacin-treated biofilms was 2.2·103 CFU/coupon, but it increased to 2.3·104 CFU/coupons when ciprofloxacin was applied with the galactan. This effect was even more pronounced in the case of amikacin. When biofilms were treated with amikacin the number of viable cells was often lower than the detection limit (~40 CFU/coupon). When the galactan was added, this number reached 2.0·106 CFU/coupon with the average value of 3.2·104 CFU/coupon.
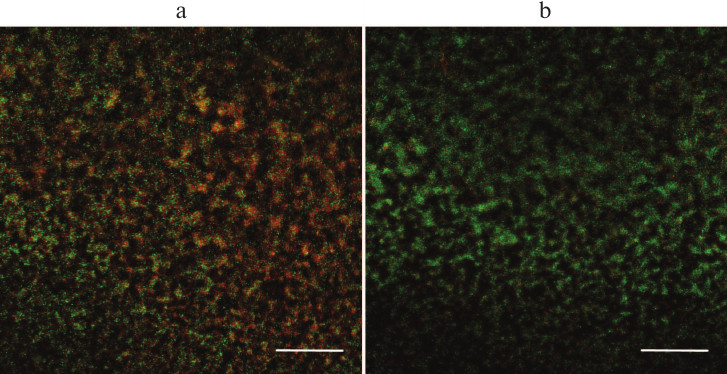
This observed effect could be explained in two ways. On one hand, the galactan might increase the antibiotic tolerance of the bacterial cells. Alternatively, it is conceivable that a portion of bacterial cells is not killed, but enters non-culturable state upon antibiotic treatment, and the galactan prevents this transition, leading to the perceived increase in the viable cell counts. To determine which of the two explanations is more probable, PAO1 biofilms treated with amikacin or amikacin in combination with the galactan were studied by fluorescence microscopy (Fig. 4). To discriminate between viable and dead cells, the biofilms were stained with LIVE/DEAD fluorescent dyes. These dyes stain viable bacteria green, while dead cells and extracellular DNA are stained red. Figure 4 shows that a biofilm treated with amikacin in combination with potato galactan contains many more viable cells compared to amikacin-only treated biofilm. Apparently, the galactan somehow protects PAO1 biofilms from antibiotics. Interestingly, this effect was not observed when planktonic cells were incubated with the galactan and antibiotics at the same concentrations.
Fig. 4. Biofilms of P. aeruginosa PAO1 treated with amikacin (a) or amikacin with potato galactan (b) and stained with fluorescent LIVE/DEAD dyes. The scale bar is 100 μm.
Next, we decided to test if the galactan exerts a similar effect on other P. aeruginosa strains, specifically clinical isolate 216 that was used in the previous work [22]. We chose to test the combined effect of amikacin with the galactan against isolate 216 biofilms since this antibiotic demonstrated a more pronounced effect against PAO1 biofilms. The MIC and MBC of amikacin against isolate 216 were the same as for PAO1, 32 and 64 μg/ml for the initial bacterial density of 5·105 CFU/ml, and 64 and 256 μg/ml for the inoculum of 5·107 CFU/ml (table). The biofilms of isolate 216 were somewhat more resistant to amikacin, having MBEC of 512 μg/ml (table). The galactan had no effect on amikacin MBEC against isolate 216, similarly to what was observed with PAO1. Likewise, when isolate 216 biofilms were cultivated on polypropylene coupons and treated with amikacin, the addition of 1 mg/ml galactan lead to a significant increase in the viable cell counts (Fig. 3).
Potato galactan-induced biofilm dispersion stimulates renewal of bacterial growth after amikacin treatment. PAO1 biofilms treated with amikacin at a concentration of 256 μg/ml (MBEC) could not resume bacterial growth in fresh nutrient broth. However, the fluorescence microscopy demonstrated the presence of viable bacteria in such biofilms (Fig. 4a). Apparently, these bacterial cells had entered a non-culturable state, which led to the lack of bacterial growth after amikacin treatment.
Nevertheless, biofilms cultivated on polypropylene coupons and treated with 256 μg/ml amikacin could still resume growth in fresh nutrient broth if they had been disrupted by glass beads. In other words, the determined value of amikacin MBEC was not the true minimum concentration required for biofilm eradication. In this case, the biofilm eradication was only apparent and resulted from the transition of surviving bacteria to the non-culturable state. Importantly, not only physical disruption but treatment with 1 mg/ml galactan as well stimulated the renewal of bacterial growth under the conditions when the unperturbed biofilms stayed in the non-culturable state. This effect could not influence the viable cell counts in amikacin-treated biofilms, since the biofilms were physically disrupted before plate counting. Besides, the fluorescence microscopy results indicate that potato galactan indeed protects biofilm-dwelling bacteria from antibiotics. Nevertheless, the observation that amikacin-treated biofilms resume their growth after physical disruption and the galactan treatment alike serves as additional evidence of the influence of the galactan on P. aeruginosa biofilm structure.
DISCUSSION
In the present work, we demonstrated the ability of potato galactan to effectively suppress P. aeruginosa biofilm formation on various surfaces and to partially disperse pre-formed biofilms. However, the galactan failed to enhance antibiotic efficacy during combined treatment. On the contrary, the viable cell counts in biofilms treated with an antibiotic in combination with the galactan were much higher compared to biofilms treated with the antibiotic alone.
Potato galactan at a concentration of 1 mg/ml effectively prevented P. aeruginosa biofilm formation on various substrates. We demonstrated previously that the galactan inhibited the early stages (5 h) of biofilm formation by P. aeruginosa clinical isolate 216. In this work, the biofilm inhibition effect was reproduced for laboratory strain PAO1 using longer cultivation times (24 h). Moreover, the galactan partially dispersed pre-formed PAO1 biofilms. Apparently, the galactan does not prevent the initial bacterial adhesion but interferes with the microcolony formation and the biomass build-up (Fig. 2). We suppose that the biofilm inhibition effect of the galactan results from excessive or abnormal aggregation of bacterial cells by the galactan, which prevents the formation of proper biofilm architecture. Indeed, large aggregates were observed in the planktonic fraction when biofilms grown in presence of the galactan were washed before analysis. Excessive aggregation as the mechanism of biofilm inhibition is further supported by the fact that lower galactan concentrations stimulated biomass accumulation in the biofilms. Apparently, lower galactan concentrations cause lower levels of aggregation that are compatible with normal biofilm development. Such a mechanism of action has not been reported for polysaccharides before. In those works where the authors studied the influence of polysaccharides on bacterial aggregation, antibiofilm polysaccharides always prevented aggregation [7, 8, 27, 28]. However, a similar effect was described for Bordetella holmesii BipA protein [32]. Bordetella holmesii isolates defective in the bipA gene form large aggregates during cultivation and do not form biofilms. When a functional copy of the bipA gene is present, aggregation is suppressed, allowing normal biofilm formation by B. holmesii [32]. Thus, although BipA, unlike the galactan, prevents aggregation instead of stimulating it, the general mechanism of biofilm inhibition due to excessive bacterial aggregation can be similar in these cases.
On the molecular level, a possible receptor of galactan could be LecA. LecA is a secreted lectin that binds monomeric galactose and galactose residues in oligo- and polysaccharides. Along with LecB that binds mannose and fucose, LecA is involved in P. aeruginosa biofilm formation, bacterial adhesion to epithelial cells, and is a potential virulence factor [33, 34]. Both lectins are targets for the development of antibiofilm compounds [34, 35]. Earlier we demonstrated that galactan inhibited LecA-induced hemagglutination [22]. Interestingly, one of the LecA inhibitors described in the literature, namely galactosylated calix[4]arene-based glycocluster, caused aggregation of P. aeruginosa planktonic cells when applied in a certain concentration range, and this effect was not observed with a lecA-deficient mutant [36]. This compound also inhibited PAO1 biofilm formation. However, the latter effect was probably caused by a mechanism other than LecA inhibition [36]. On the other hand, our preliminary experiments indicated that potato galactan aggregated Stenotrophomonas maltophilia planktonic cells and modified its biofilm structure (data not shown). However, LecA has not been identified in this species so far.
Despite the ability of the galactan to inhibit biofilm formation and partially disperse pre-formed biofilms, this polysaccharide protected biofilm-dwelling bacterial cells when applied simultaneously with antibiotics. The mechanism of this protection is unclear. Since this effect does not extend to planktonic cells, the most probable explanation is the partial remodeling of biofilm architecture by the galactan. This remodeling could impede antibiotic penetration to bacterial cells or stimulate the alterations in their physiological state, which in turn could lead to increased antibiotic tolerance. It must be noted that a similar effect towards P. aeruginosa biofilms was recently described for Staphylococcus aureus protein A (SpA) [37]. SpA was shown to interact with biofilm matrix polysaccharide psl and type IV pili [38]. When SpA was added to pre-formed biofilms, these interactions changed the normal biofilm structure and led to the formation of large and dense bacterial aggregates. Importantly, this remodeling of biofilm architecture increased its tolerance to tobramycin, an aminoglycoside antibiotic similar to amikacin [37]. Although the mechanisms of biofilm remodeling by potato galactan and SpA are probably different, it can be speculated that this remodeling leads to a denser aggregation of bacterial cells and increased antibiotic tolerance in both cases.
In the present work, we studied the effect of potato galactan on P. aeruginosa biofilms, including the effect of galactan treatment in combination with antibiotics ciprofloxacin and amikacin. The galactan inhibited the formation of P. aeruginosa biofilms and partially dispersed pre-formed biofilms. This effect is presumably caused by excessive aggregation of bacterial cells due to the interaction with the galactan. However, the galactan did not enhance the antibiotic efficacy towards P. aeruginosa biofilms during combined treatment. On the contrary, the galactan protected the biofilm-dwelling bacteria from antibiotics. This is apparently connected to galactan-induced remodeling of biofilm architecture. Similar effect towards P. aeruginosa biofilms was earlier demonstrated for S. aureus protein A. It is unlikely that potato galactan can be used as the basis for further development of antibiofilm drugs. Nevertheless, our results appear interesting and demonstrate a tight connection between biofilm structure and its antibiotic susceptibility.
Acknowledgements
The authors are thankful to Dr. I. G. Tiganova for P. aeruginosa PAO1 strain, Dr. O. Yu. Dobrynina for P. aeruginosa isolate 216, and Dr. Yu. M. Romanova for kindly providing LIVE/DEAD dyes.
Funding
This work was supported by the Ministry of Healthcare of the Russian Federation in the frame of government assignment no. NIOKR 115030470038.
Conflict of Interest
The authors declare no conflict of interest in financial or any other sphere.
Ethical Approval
This article does not contain any studies with human participants or animals.
REFERENCES
1.Hall-Stoodley, L., Costerton, J. W., and Stoodley,
P. (2004) Bacterial biofilms: from the natural environment to
infectious diseases, Nat. Rev. Microbiol., 2, 95-108;
doi: 10.1038/nrmicro821.
2.Hall-Stoodley, L., and Stoodley, P. (2009) Evolving
concepts in biofilm infections, Cell Microbiol., 11,
1034-1043; doi: 10.1111/j.1462-5822.2009.01323.x.
3.Hoiby, N., Bjarnsholt, T., Givskov, M., Molin, S.,
and Ciofu, O. (2010) Antibiotic resistance of bacterial biofilms,
Int. J. Antimicrob. Agents., 35, 322-332; doi:
10.1016/j.ijantimicag.2009.12.011.
4.Jensen, P. O., Givskov, M., Bjarnsholt, T., and
Moser, C. (2010) The immune system vs. Pseudomonas aeruginosa
biofilms, FEMS Immunol. Med. Microbiol., 59, 292-305;
doi: 10.1111/j.1574-695X.2010.00706.x.
5.Mah, T.-F. (2010) Biofilm-specific antibiotic
resistance, Future Microbiol., 7, 1061-1072; doi:
10.2217/fmb.12.76.
6Costerton, J. W. (1999) Bacterial biofilms: a common cause of
persistent infections, Science, 284, 1318-1322; doi:
10.1126/science.284.5418.1318.
7.Valle, J., Da Re, S., Henry, N., Fontaine, T.,
Balestrino, D., Latour-Lambert, P., and Ghigo, J.-M. (2006)
Broad-spectrum biofilm inhibition by a secreted bacterial
polysaccharide, Proc. Natl. Acad. Sci. USA, 103,
12558-12563; doi: 10.1073/pnas.0605399103.
8.Jiang, P., Li, J., Han, F., Duan, G., Lu, X., Gu,
Y., and Yu, W. (2011) Antibiofilm activity of an exopolysaccharide from
marine bacterium Vibrio sp. QY101, PLoS One, 6,
e18514; doi: 10.1371/journal.pone.0018514.
9.Kanmani, P., Satish Kumar, R., Yuvaraj, N., Paari,
K. A., Pattukumar, V., and Arul, V. (2011) Production and purification
of a novel exopolysaccharide from lactic acid bacterium
Streptococcus phocae PI80 and its functional characteristics
activity in vitro, Bioresour. Technol., 102,
4827-4833; doi: 10.1016/j.biortech.2010.12.118.
10.Sayem, S. M. A., Manzo, E., Ciavatta, L.,
Tramice, A., Cordone, A., Zanfardino, A., De Felice, M., and
Varcamonti, M. (2011) Anti-biofilm activity of an exopolysaccharide
from a sponge-associated strain of Bacillus licheniformis,
Microb. Cell. Fact., 10, 74; doi:
10.1186/1475-2859-10-74.
11.Spano, A., Lagana, P., Visalli, G., Maugeri, T.
L., and Gugliandolo, C. (2016) In vitro antibiofilm activity of
an exopolysaccharide from the marine thermophilic Bacillus
licheniformis T14, Curr. Microbiol., 72, 518-528;
doi: 10.1007/s00284-015-0981-9.
12.Wu, S., Liu, G., Jin, W., Xiu, P., and Sun, C.
(2016) Antibiofilm and anti-infection of a marine bacterial
exopolysaccharide against Pseudomonas aeruginosa, Front.
Microbiol., 7, 102; doi: 10.3389/fmicb.2016.00102.
13.Brian-Jaisson, F., Molmeret, M., Fahs, A.,
Guentas-Dombrowsky, L., Culioli, G., Blache, Y., Cerantola, S., and
Ortalo-Magne, A. (2016) Characterization and anti-biofilm activity of
extracellular polymeric substances produced by the marine
biofilm-forming bacterium Pseudoalteromonas ulvae strain TC14,
Biofouling, 32, 547-560; doi:
10.1080/08927014.2016.1164845.
14.Papa, R., Parrilli, E., Sannino, F., Barbato, G.,
Tutino, M. L., Artini, M., and Selan, L. (2013) Anti-biofilm activity
of the Antarctic marine bacterium Pseudoalteromonas haloplanktis
TAC125, Res. Microbiol., 164, 450-456; doi:
10.1016/j.resmic.2013.01.010.
15.Kanmani, P., Suganya, K., Satish Kumar, R.,
Yuvaraj, N., Pattukumar, V., Paari, K. A., and Arul, V. (2013)
Synthesis and functional characterization of antibiofilm
exopolysaccharide produced by Enterococcus faecium MC13 isolated
from the gut of fish, Appl. Biochem. Biotechnol., 169,
1001-1015; doi: 10.1007/s12010-012-0074-1.
16.Kavita, K., Singh, V. K., Mishra, A., and Jha, B.
(2014) Characterization and anti-biofilm activity of extracellular
polymeric substances from Oceanobacillus iheyensis,
Carbohydr. Polym., 101, 29-35; doi:
10.1016/j.carbpol.2013.08.099.
17.Li, Y., Li, Q., Hao, D., Jiang, D., Luo, Y., Liu,
Y., and Zhao, Z. (2015) Production, purification, and antibiofilm
activity of a novel exopolysaccharide from Arthrobacter sp. B4,
Prep. Biochem. Biotechnol., 45, 192-204; doi:
10.1080/10826068.2014.907180.
18.Pradeepa, Shetty, A. D., Matthews, K., Hegde, A.
R., Akshatha, B., Mathias, A. B., Mutalik, S., and Vidya, S. M. (2016)
Multidrug resistant pathogenic bacterial biofilm inhibition by
Lactobacillus plantarum exopolysaccharide, Bioact. Carbohydr.
Diet Fibre, 8, 7-14; doi: 10.1016/j.bcdf.2016.06.002.
19.Rendueles, O., Travier, L., and Latour-Lambert,
P. (2011) Screening of Escherichia coli species biodiversity
reveals new biofilm-associated antiadhesion polysaccharide,
MBio, 2, e00043-11; doi: 10.1128/mBio.00043-11.
20.Bendaoud, M., Vinogradov, E., Balashova, N. V.,
Kadouri, D. E., Kachlany, S. C., and Kaplan, J. B. (2011)
Broad-spectrum biofilm inhibition by Kingella kingae
exopolysaccharide, J. Bacteriol., 193, 3879-3886; doi:
10.1128/JB.00311-11.
21.Dos Santos Goncalves, M., Delattre, C.,
Balestrino, D., Charbonnel, N., Elboutachfaiti, R., Wadouachi, A.,
Badel, S., Bernardi, T., Michaud, P., and Forestier, C. (2014)
Anti-biofilm activity: a function of Klebsiella pneumoniae
capsular polysaccharide, PLoS One, 9, e99995; doi:
10.1371/journal.pone.0099995.
22.Grishin, A., Karyagina, A. S., Tiganova, I. G.,
Dobrynina, O. Y., Bolshakova, T. N., Boksha, I. S., Alexeyeva, N. V.,
Stepanova, T. V., Lunin, V. G., Chuchalin, A. G., and Ginzburg, A. L.
(2013) Inhibition of Pseudomonas aeruginosa biofilm formation by
LecA-binding polysaccharides, Int. J. Antimicrob. Agents,
42, 471-472; doi: 10.1016/j.ijantimicag.2013.07.003.
23.Vazquez-Rodriguez, A., Vasto-Anzaldo, X. G.,
Barboza Perez, D., Vázquez-Garza, E., Chapoy-Villanueva, H.,
Garcia-Rivas, G., Garza-Cervantes, J., Gomez-Lugo, J. J., Gomez-Loredo,
A. E., Gonzalez, M. T. G., Zarate, X., and Morones-Ramirez, J. R.
(2018) Microbial competition of Rhodotorula mucilaginosa
UANL-001L and E. coli increase biosynthesis of non-toxic
exopolysaccharide with applications as a wide-spectrum antimicrobial,
Sci. Rep., 8, 798; doi: 10.1038/s41598-017-17908-8.
24.Doghri, I., Lavaud, J., Dufour, A., Bazire, A.,
Lanneluc, I., and Sable, S. (2017) Cell-bound exopolysaccharides from
an axenic culture of the intertidal mudflat Navicula phyllepta
diatom affect biofilm formation by benthic bacteria, J. Appl.
Phycol., 29, 165-177; doi: 10.1007/s10811-016-0943-z.
25.Bernal, P., and Llamas, M. A. (2012) Promising
biotechnological applications of antibiofilm exopolysaccharides,
Microb. Biotechnol., 5, 670-673; doi:
10.1111/j.1751-7915.2012.00359.x.
26.Rendueles, O., Kaplan, J. B., and Ghigo, J.-M.
(2013) Antibiofilm polysaccharides, Environ. Microbiol.,
15, 334-346; doi: 10.1111/j.1462-2920.2012.02810.x.
27.Kim, Y., Oh, S., and Kim, S. H. (2009) Released
exopolysaccharide (r-EPS) produced from probiotic bacteria reduce
biofilm formation of enterohemorrhagic Escherichia coli O157:H7,
Biochem. Biophys. Res. Commun., 379, 324-329; doi:
10.1016/j.bbrc.2008.12.053.
28.Karwacki, M. T., Kadouri, D. E., Bendaoud, M.,
Izano, E. A., Sampathkumar, V., Inzana, T. J., and Kaplan, J. B. (2013)
Antibiofilm activity of Actinobacillus pleuropneumoniae serotype
5 capsular polysaccharide, PLoS One, 8, e63844; doi:
10.1371/journal.pone.0063844.
29.O’Toole, G. A. (2011) Microtiter dish
biofilm formation assay, J. Vis. Exp., 47, 2437; doi:
10.3791/2437.
30.Mah, T.-F. (2014) Establishing the minimal
bactericidal concentration of an antimicrobial agent for planktonic
cells (MBC-P) and biofilm cells (MBC-B), J. Vis. Exp.,
83, e50854; doi: 10.3791/50854.
31.Schindelin, J., Arganda-Carreras, I., Frise, E.,
Kaynig, V., Longair, M., Pietzsch, T., Preibisch, S., Rueden, C.,
Saalfeld, S., Schmid, B., Tinevez, J.-Y., White, D. J., Hartenstein,
V., Eliceiri, K., Tomancak, P., and Cardona, A. (2012) Fiji: an
open-source platform for biological-image analysis, Nat. Meth.,
9, 676-682; doi: 10.1038/nmeth.2019.
32.Hiramatsu, Y., Saito, M., Otsuka, N., Suzuki, E.,
Watanabe, M., Shibayama, K., and Kamachi, K. (2016) BipA is associated
with preventing autoagglutination and promoting biofilm formation in
Bordetella holmesii, PLoS One, 11, e0159999; doi:
10.1371/journal.pone.0159999.
33.Imberty, A., Wimmerova, M., Mitchell, E. P., and
Gilboa-Garber, N. (2004) Structures of the lectins from Pseudomonas
aeruginosa: insights into the molecular basis for host glycan
recognition, Microbes Infect., 6, 221-228; doi:
10.1016/j.micinf.2003.10.016.
34.Grishin, A. V., Krivozubov, M. S., Karyagina, A.
S., and Gintsburg, A. L. (2015) Pseudomonas aeruginosa lectins
as targets for novel antibacterials, Acta Naturae, 7,
29-41.
35.Titz, A. (2014) Carbohydrate-based anti-virulence
compounds against chronic Pseudomonas aeruginosa infections with
a focus on small molecules, in Carbohydrates as Drugs. Topics in
Medicinal Chemistry, Vol. 12 (Seeberger, P., and Rademacher, C.,
eds.) Springer, Cambridge, pp. 169-186; doi: 10.1007/7355_2014_44.
36.Boukerb, A. M., Rousset, A., Galanos, N., Mear,
J.-B., Thepaut, M., Grandjean, T., Gillon, E., Cecioni, S.,
Abderrahmen, C., Faure, K., Redelberger, D., Kipnis, E., Dessein, R.,
Havet, S., Darblade, B., Matthews, S. E., de Bentzmann, S., Guery, B.,
Cournoyer, B., Imberty, A., and Vidal, S. (2014) Antiadhesive
properties of glycoclusters against Pseudomonas aeruginosa lung
infection, J. Med. Chem., 57, 10275-10289; doi:
10.1021/jm500038p.
37.Beaudoin, T., Yau, Y. C. W., Stapleton, P. J.,
Gong, Y., Wang, P. W., Guttman, D. S., and Water, V. (2017)
Staphylococcus aureus interaction with Pseudomonas
aeruginosa biofilm enhances tobramycin resistance, NPJ Biofilms
Microbiomes, 3, 25; doi: 10.1038/s41522-017-0035-0.
38.Armbruster, C. R., Wolter, D. J., Mishra, M.,
Hayden, H. S., Radey, M. C., Merrihew, G., Maccoss, M. J., Burns, J.,
Wozniak, D. J., Parsek, M. R., and Hoffman, L. R. (2016)
Staphylococcus aureus protein A mediates interspecies
interactions at the cell surface of Pseudomonas aeruginosa,
MBio, 7, e00538-16; doi: 10.1128/mBio.00538-16.
Supplementary Material (MS Excel)